library(dplyr)
library(tidyverse)
library(spdep)
library(spData)
library(spgwr)
library(mgwrsar)
library(rgdal)
library(raster)
library(DataExplorer)
library(tigris)
library(sf)
library(Rcpp)
library(ggplot2)
library(corrplot)
library(writexl)
library(nortest)
library(car)
library(DescTools)
library(lmtest)
library(tseries)3. MGWR-SAR
Deskripsi
SAR yang juga dikenal sebagai spatial lag adalah analisis regresi spasial dengan efek dependensi spasial pada peubah respon. Menurut Anselin (1988), model SAR merupakan model regresi spasial yang dapat memperhitungkan efek dependensi spasial. Namun, Shekhar et al. (2018) menyatakan bahwa model SAR tidak cocok digunakan pada kasus heterogenitas spasial, sehingga perlu dilakukan pemodelan dengan regresi terboboti geografis (Geographically Weighted Regression/GWR). Pendekatan GWR merupakan modifikasi dari model regresi klasik yang dapat memperhitungkan efek heterogenitas spasial. Model GWR menghasilkan penduga parameter lokal untuk setiap lokasi pengamatan dengan setiap parameter dihitung pada setiap titik lokasi geografis (Lu et al., 2014). Dalam analisisnya, GWR menggunakan matriks pembobot yang tergantung pada kedekatan lokasi dengan fungsi kernel.
Faktanya, tidak semua parameter regresi dalam model GWR bervariasi secara spasial, beberapa parameter mungkin tidak signifikan secara spasial. Model GWR kemudian dikembangkan menjadi model Mixed Geographically Weighted Regression (MGWR) yang menggabungkan model regresi linier dan GWR. Dalam MGWR, beberapa parameter GWR tetap konstan di semua lokasi, sementara yang lain bervariasi pada lokasi pengamatan. Hal ini menghasilkan penduga parameter global dan lokal dalam model MGWR (Fotheringham et al., 2002).
Pada suatu data kadang mengandung efek dependensi dan keragaman spasial. Geniaux & Martinetti (2018) mengembangkan MGWR dengan penambahan model dependensi SAR yang dikenal dengan Mixed Geographically Weighted Regression-Spatial Autoregressive (MGWR-SAR). Beberapa model yang dikembangkan diberi notasi MGWR-SAR \((0, k_c, k_v)\), dan MGWR-SAR \((1, k_c, k_v)\). MGWR-SAR \((0, k_c, k_v)\) menunjukkan bahwa model mempunyai koefisien dependensi (\(\rho\)) yang sama di semua lokasi, sedangkan MGWR-SAR \((1, k_c,k_v)\) menunjukkan koefisien dependensi (\(\rho\)) yang berbeda-beda antar lokasi. Konstanta \(k_c\) menunjukkan banyaknya parameter regresi global dan \(k_v\) menunjukkan banyaknya parameter regresi lokal.
MGWR-SAR memiliki notasi \((i_{\rho},k_c,k_v)\) dimana \(i_{\lambda}\) dapat dinyatakan bahwa lag konstan (\(\rho=0\)) atau bervariasi secara spasial (\(\rho=1\)), \(k_c\) merupakan jumlah parameter regresi global, dan \(k_v\) adalah jumlah parameter regresi lokal. Model MGWR-SAR \((0, k_c,k_v)\) dan MGWR-SAR \((1, k_c, k_v)\) dituliskan sebagai berikut:
\[ y_i = \rho W y_i + \beta_0 (u_i, v_i) + \sum^q_{k=1} \beta_k (u_i, v_i)X_{ik} + \sum^p_{k=q+1} \beta_k X_{ik} + \epsilon_i \quad (MGWR{\text -}SAR(0,k_c,k_v)) \tag{1}\]
\[ y_i = \rho (u_i, v_i) W y_i + \beta_0 (u_i, v_i) + \sum^q_{k=1} \beta_k (u_i, v_i)X_{ik} + \sum^p_{k=q+1} \beta_k X_{ik} + \epsilon_i \quad (MGWR{\text -}SAR(1,k_c,k_v)) \tag{2}\]
\(W y_i\) adalah matriks bobot spasial yang menghubungkan nilai peubah respon \(y_i\) di lokasi \((u_i,v_i)\).
MGWR-SAR pada studi kasus HIV
Data
Penelitian ini mengacu pada Djuraidah et al. (2023). Data yang digunakan merupakan data sekunder hasil pemantauan perkembangan situasi HIV tahun 2018 dan peubah-peubah yang diduga berperan dalam memengaruhi peubah respon. Data diperoleh dari Kementerian Kesehatan, Badan Perencanaan Pembangunan Nasional (Bappenas), dan Badan Pusat Statistik (BPS). Jumlah lokasi amatan dengan total kasus HIV tidak nol sebanyak 390 kabupaten/kota dari seluruh Indonesia.
Peubah yang dijadikan acuan dalam penelitian ini yaitu kasus HIV per 100.000 penduduk sebagai peubah respon. Populasi kunci per 100.000 penduduk, kasus positif pada ibu hamil per 100.000 penduduk, pasien tuberkulosis per 100.000 penduduk, tingkat kemiskinan, dan tingkat pengangguran sebagai peubah prediktor. Metode yang digunakan adalah MGWR, MGWR-SAR \((0, k_c,k_v)\), dan MGWR-SAR \((1, k_c,k_v)\).
Tahapan Analisis Data
Berikut adalah urutan langkah-langkah yang dilakukan dalam penelitian ini:
Melakukan eksplorasi data untuk mengetahui pola sebaran peubah respon dan plot pencaran antara peubah respon dan prediktor.
Melakukan uji multikolinieritas dengan menggunakan Variance Inflation Factor (VIF).
Menguji efek spasial dengan menerapkan uji dependensi spasial pada respon dan galat menggunakan uji Robust Lagrange Multiplier (RLM) (Anselin, 1988) dan menguji efek keragaman spasial melalui uji Breusch-Pagan (Arbia, 2006).
Menetapkan matriks pembobot dengan membandingkan empat fungsi pembobot yang berbeda, yaitu Fixed Gaussian, Fixed Bisquare, Adaptive Gaussian, dan Adaptive Bisquare. Fungsi pembobot yang optimal ditentukan berdasarkan nilai Akaike Information Criterion (AIC) terkecil.
Melakukan pemodelan GWR dengan fungsi pembobot optimal untuk mengidentifikasi parameter yang berpengaruh secara global maupun lokal.
Menduga parameter dalam model MGWR dengan fungsi pembobot optimal.
Melakukan pendugaan parameter model MGWR-SAR \((0, k_c,k_v)\) dan MGWR-SAR \((1, k_c,k_v)\).
Membandingkan hasil model MGWR, MGWR-SAR \((0, k_c,k_v)\), dan MGWR-SAR \((1, k_c,k_v)\).
Menerapkan uji Wald untuk menguji signifikansi parameter.
Melakukan interpretasi pada hasil analisis.
MGWR-SAR pada kajian Sosial Ekonomi
Data
Data dalam penelitian ini berasal dari data sekunder Badan Pusat Statistik (BPS) mengenai aspek sosial ekonomi masyarakat dengan objek amatan 514 Kabupaten/Kota seluruh Indonesia pada tahun 2021. Berikut adalah peubah-peubah yang digunakan (Tabel 1):
| Peubah | Keterangan | Satuan |
|---|---|---|
| \(Y\) | PDRB atas Dasar Harga Konstan menurut Pengeluaran | Rupiah |
| \(X_1\) | Persentase Penduduk Miskin (P0) Menurut Kabupaten/Kota | Persen |
| \(X_2\) | Rata-rata Lama Sekolah Penduduk usia > 15 tahun | Tahun |
| \(X_3\) | Pengeluaran per Kapita Disesuaikan | Ribu Rupiah/Orang/Tahun |
| \(X_4\) | Indeks Pembangunan Manusia | - |
| \(X_5\) | Umur Harapan Hidup | Tahun |
| \(X_6\) | Persentase Rumah Tangga yang Memiliki Akses terhadap Sanitasi Layak | Persen |
| \(X_7\) | Persentase Rumah Tangga yang Memiliki Akses terhadap Air Minum Layak | Persen |
| \(X_8\) | Tingkat Pengangguran Terbuka | Persen |
| \(X_9\) | Tingkat Partisipasi Angkatan Kerja | Persen |
Metode Analisis
Langkah-langkah yang dilakukan untuk menganalisis data adalah sebagai berikut:
Melakukan eksplorasi data untuk melihat karakteristik data serta membuat visualisasi peta tematik pada setiap peubah.
-
Model Regresi Linear Berganda (RLB)
- Membangun model RLB
- Mengidentifikasi adanya multikolinearitas melalui Variance Inflation Factor (VIF)
- Diagnostik sisaan model RLB meliputi kenormalan, kehomogenan ragam, dan kebebasan sisaan.
-
Model Dependensi Spasial
- Membentuk matriks pembobot spasial berbasis ketetanggaan dan jarak yang dinormalisasi baris, antara lain rook contiguity, queen contiguity, k-nearest neighbor (KNN), inverse distance weight (IDW), dan eksponensial.
- Mengecek adanya autokorelasi spasial dengan menggunakan:
- Indeks Moran
-
Hipotesis:
[H_0 : I = 0 ()] [H_1 : I ()]
-
Statistik Uji:
\(Z(I) = \frac{I - E(I)}{\sqrt{\text{Var}(I)}}\)
\(I\) adalah Statistik Indeks Moran (\(I > 0\) terdapat autokorelasi positif; \(I < 0\) terdapat autokorelasi negatif)
Kriteria keputusan jika \(Z(I) > Z_{\alpha/2}\)
-
- Indeks Moran
- Menyelidiki adanya dependensi spasial dengan Uji Lagrange Multiplier.
- Menyelidiki adanya keragaman spasial dengan menggunakan uji Breusch-Pagan.
- Membangun model dependensi spasial yang sesuai dan uji parameter.
- Diagnostik sisaan model.
-
Model Dependensi dan Keragaman Spasial
- Menentukan matriks pembobot kernel.
- Membentuk bandwidth optimum.
- Membangun model keragaman dan dependensi spasial, antara lain:
- Model GWR
- Model MGWR−SAR \((0, k, 0)\)
- Model MGWR−SAR \((0, 0, k)\)
- Membandingkan kebaikan model dengan kriteria RMSE.
- Visualisasi dugaan parameter model terbaik.
Tahapan Analisis dengan R
Package
Load Data
pdrb <- read.csv("data/pdrb.csv", sep = ";")
glimpse(pdrb)
#> Rows: 514
#> Columns: 13
#> $ Nama.Wilayah <chr> "Simeulue", "Aceh Singkil", "Aceh Selatan", "…
#> $ latitude <dbl> 2.58, 2.36, 3.31, 3.31, 5.26, 4.45, 4.45, 5.4…
#> $ longitude <dbl> 96.1, 97.9, 97.4, 97.7, 96.0, 96.8, 96.2, 95.…
#> $ Y <int> 1648096, 1780419, 4345784, 3487157, 8433526, …
#> $ X1 <dbl> 18.98, 20.36, 13.18, 13.41, 14.45, 15.26, 18.…
#> $ X2 <dbl> 9.48, 8.68, 8.88, 9.67, 8.21, 9.86, 9.55, 10.…
#> $ X3 <int> 7148, 8776, 8180, 8030, 8577, 10780, 9593, 96…
#> $ X4 <dbl> 66.4, 69.2, 67.4, 69.4, 67.8, 73.4, 71.7, 73.…
#> $ X5 <dbl> 65.3, 67.4, 64.4, 68.2, 68.7, 68.9, 68.0, 69.…
#> $ X6 <dbl> 71.6, 69.6, 62.5, 62.7, 66.8, 90.6, 89.6, 87.…
#> $ X7 <dbl> 87.5, 78.6, 79.7, 86.7, 83.2, 90.1, 94.2, 82.…
#> $ X8 <dbl> 5.71, 8.36, 6.46, 6.43, 7.13, 2.61, 7.09, 7.7…
#> $ X9 <dbl> 71.2, 62.9, 60.9, 69.6, 59.5, 76.3, 60.0, 61.…Eksplorasi Data
pdrb %>% as.data.frame %>%
ggplot(aes(longitude, latitude)) + geom_point(aes(size=Y), color="blue", alpha=0.6) +
ggtitle("Sebaran PDRB Kabupaten Kota di Indonesia Tahun 2021") + coord_equal() + theme_bw()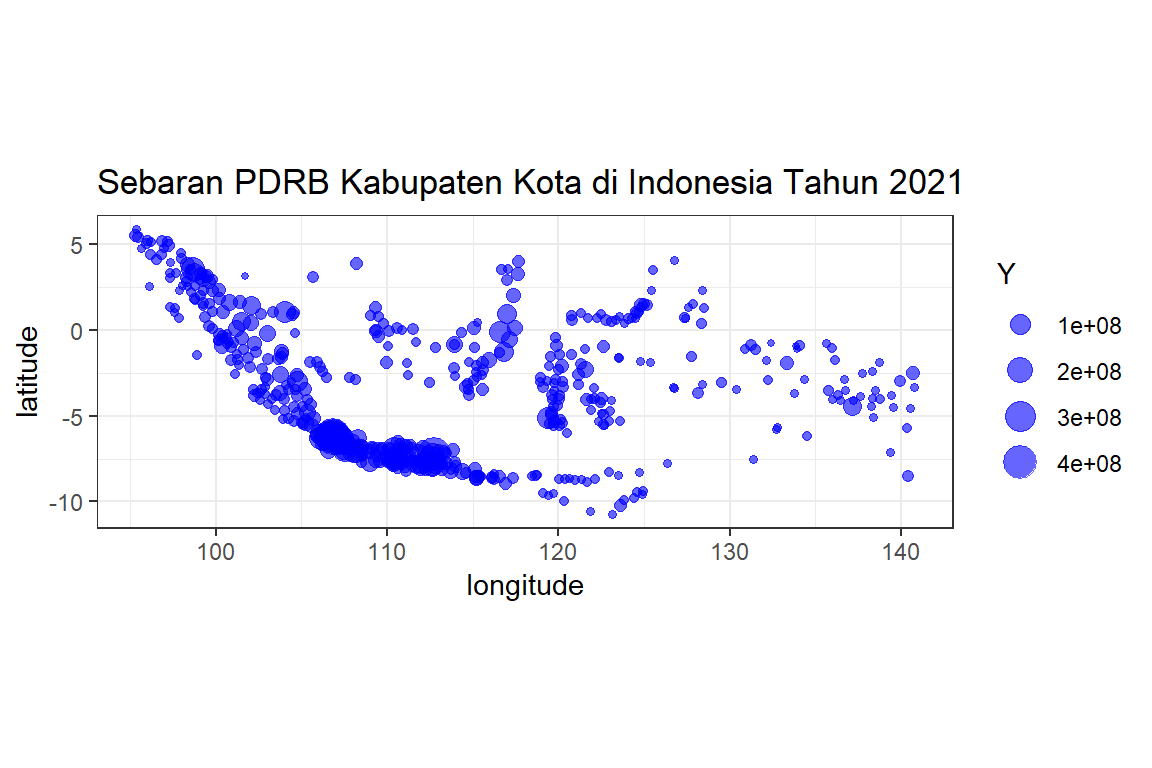
plot_intro(data = pdrb)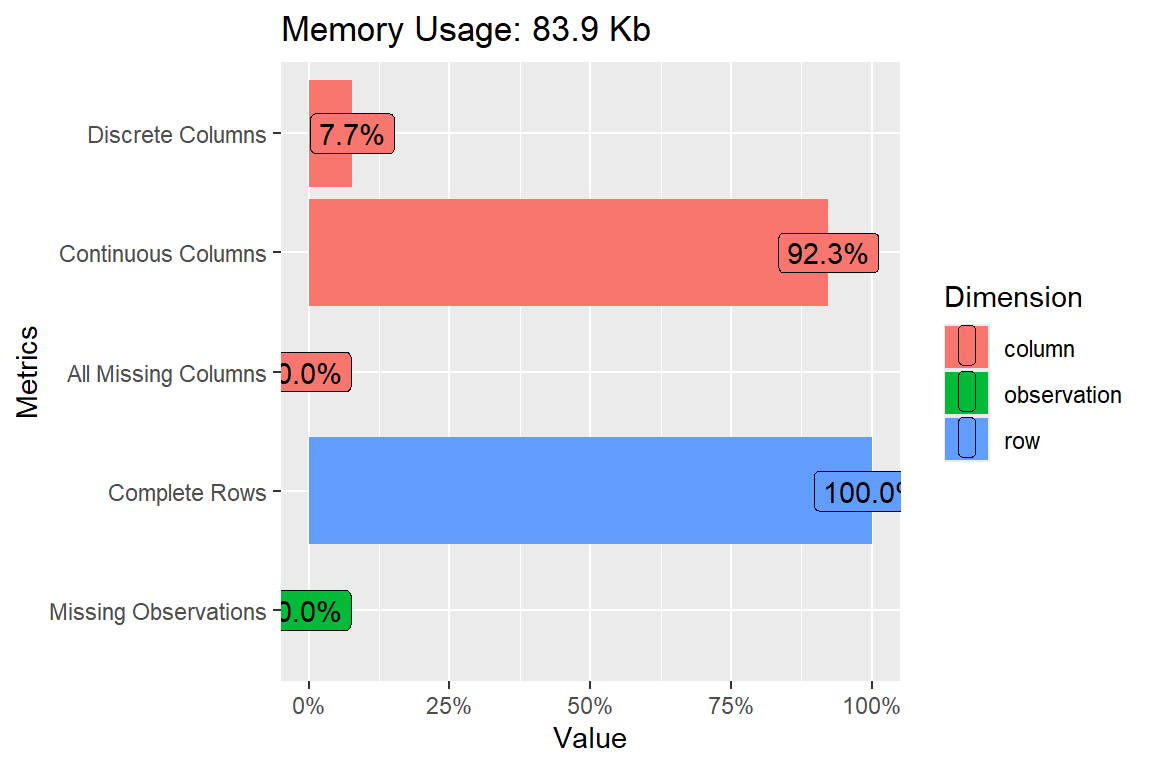
# Plot
datagwr <- pdrb[4:13]
glimpse(datagwr)
#> Rows: 514
#> Columns: 10
#> $ Y <int> 1648096, 1780419, 4345784, 3487157, 8433526, 5953118, 7…
#> $ X1 <dbl> 18.98, 20.36, 13.18, 13.41, 14.45, 15.26, 18.81, 14.05,…
#> $ X2 <dbl> 9.48, 8.68, 8.88, 9.67, 8.21, 9.86, 9.55, 10.33, 9.00, …
#> $ X3 <int> 7148, 8776, 8180, 8030, 8577, 10780, 9593, 9644, 9860, …
#> $ X4 <dbl> 66.4, 69.2, 67.4, 69.4, 67.8, 73.4, 71.7, 73.6, 70.7, 7…
#> $ X5 <dbl> 65.3, 67.4, 64.4, 68.2, 68.7, 68.9, 68.0, 69.8, 67.0, 7…
#> $ X6 <dbl> 71.6, 69.6, 62.5, 62.7, 66.8, 90.6, 89.6, 87.4, 54.1, 8…
#> $ X7 <dbl> 87.5, 78.6, 79.7, 86.7, 83.2, 90.1, 94.2, 82.4, 89.2, 9…
#> $ X8 <dbl> 5.71, 8.36, 6.46, 6.43, 7.13, 2.61, 7.09, 7.70, 7.28, 4…
#> $ X9 <dbl> 71.2, 62.9, 60.9, 69.6, 59.5, 76.3, 60.0, 61.7, 60.3, 6…
#Plot hubungan
plot_scatterplot( datagwr, by = 'Y',
ggtheme = theme_bw(),
geom_point_args = list("color" = alpha("dodgerblue4", 0.7)))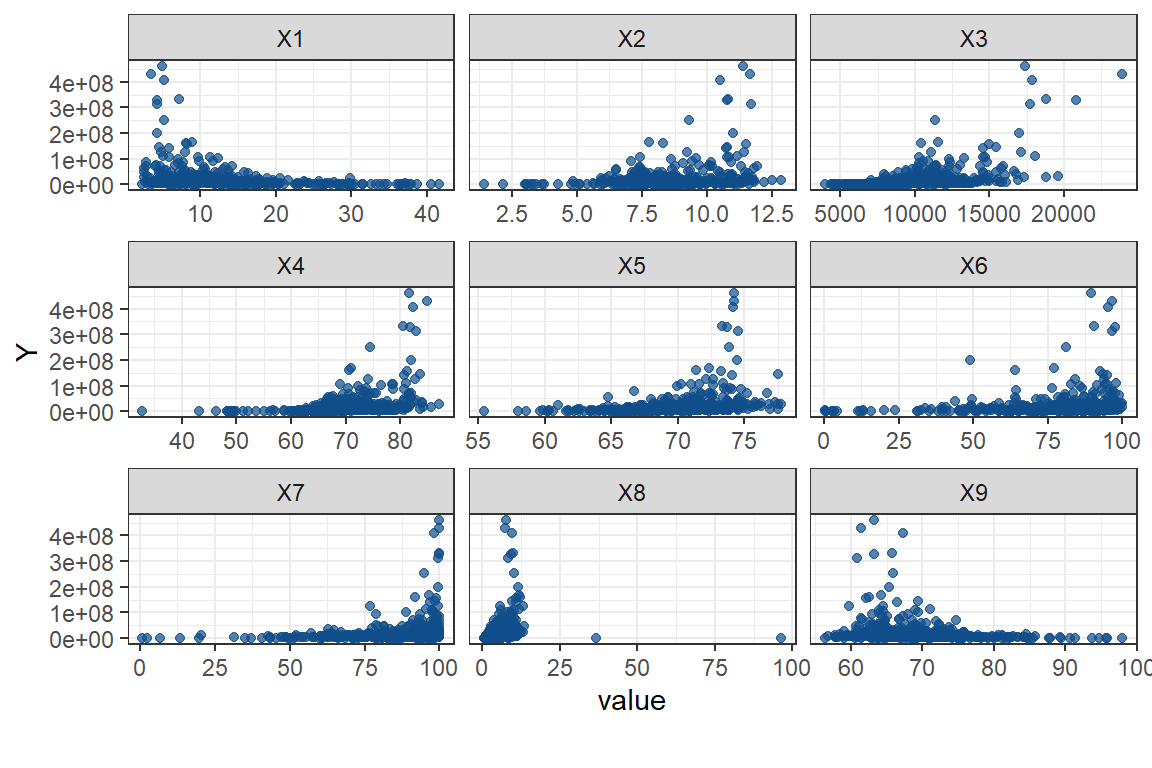
summary(datagwr)
#> Y X1 X2 X3
#> Min. :1.47e+05 Min. : 2.4 Min. : 1.42 Min. : 3976
#> 1st Qu.:3.65e+06 1st Qu.: 7.2 1st Qu.: 7.51 1st Qu.: 8574
#> Median :8.81e+06 Median :10.5 Median : 8.30 Median :10196
#> Mean :2.20e+07 Mean :12.3 Mean : 8.44 Mean :10325
#> 3rd Qu.:1.97e+07 3rd Qu.:14.9 3rd Qu.: 9.34 3rd Qu.:11719
#> Max. :4.60e+08 Max. :41.7 Max. :12.83 Max. :23888
#> X4 X5 X6 X7
#> Min. :32.8 Min. :55.4 Min. : 0.3 Min. : 0.9
#> 1st Qu.:66.6 1st Qu.:67.4 1st Qu.: 70.2 1st Qu.: 79.0
#> Median :69.6 Median :70.0 Median : 81.8 Median : 89.8
#> Mean :69.9 Mean :69.7 Mean : 77.2 Mean : 85.1
#> 3rd Qu.:73.1 3rd Qu.:72.0 3rd Qu.: 89.9 3rd Qu.: 96.4
#> Max. :87.2 Max. :77.7 Max. :100.0 Max. :100.0
#> X8 X9
#> Min. : 0.4 Min. :56.4
#> 1st Qu.: 3.2 1st Qu.:65.1
#> Median : 4.6 Median :69.0
#> Mean : 5.3 Mean :69.5
#> 3rd Qu.: 6.6 3rd Qu.:72.3
#> Max. :96.2 Max. :97.9Korelasi
#nilai p value dari korelasi antar peubah Y dan X
p.mat <- cor.mtest(datagwr)
head(p.mat$p)
#> Y X1 X2 X3 X4 X5
#> Y 0.00e+00 2.95e-08 4.04e-10 7.90e-36 2.61e-18 5.12e-14
#> X1 2.95e-08 0.00e+00 6.52e-41 1.33e-61 4.13e-79 2.47e-40
#> X2 4.04e-10 6.52e-41 0.00e+00 6.12e-68 2.04e-158 6.22e-23
#> X3 7.90e-36 1.33e-61 6.12e-68 0.00e+00 1.38e-160 5.08e-45
#> X4 2.61e-18 4.13e-79 2.04e-158 1.38e-160 0.00e+00 4.37e-79
#> X5 5.12e-14 2.47e-40 6.22e-23 5.08e-45 4.37e-79 0.00e+00
#> X6 X7 X8 X9
#> Y 2.06e-05 2.13e-07 2.45e-05 1.58e-06
#> X1 3.65e-44 2.01e-18 4.24e-01 1.38e-28
#> X2 3.04e-54 4.15e-28 1.20e-01 6.57e-34
#> X3 2.73e-45 1.56e-26 8.91e-05 2.37e-21
#> X4 6.76e-76 5.94e-42 3.15e-01 7.46e-36
#> X5 8.04e-29 5.42e-22 2.46e-01 1.00e-07
#plot korelasi
col2 <- colorRampPalette(c("#BB4444", "#EE9988", "#FFFFFF", "#77AADD", "#4477AA"))
corrplot(cor(pdrb[,4:13]), method="color", col=col2(200),
type="upper",
addCoef.col = "black",
tl.col="black",
p.mat = p.mat$p, sig.level = 0.05, insig = "blank",
diag=FALSE
)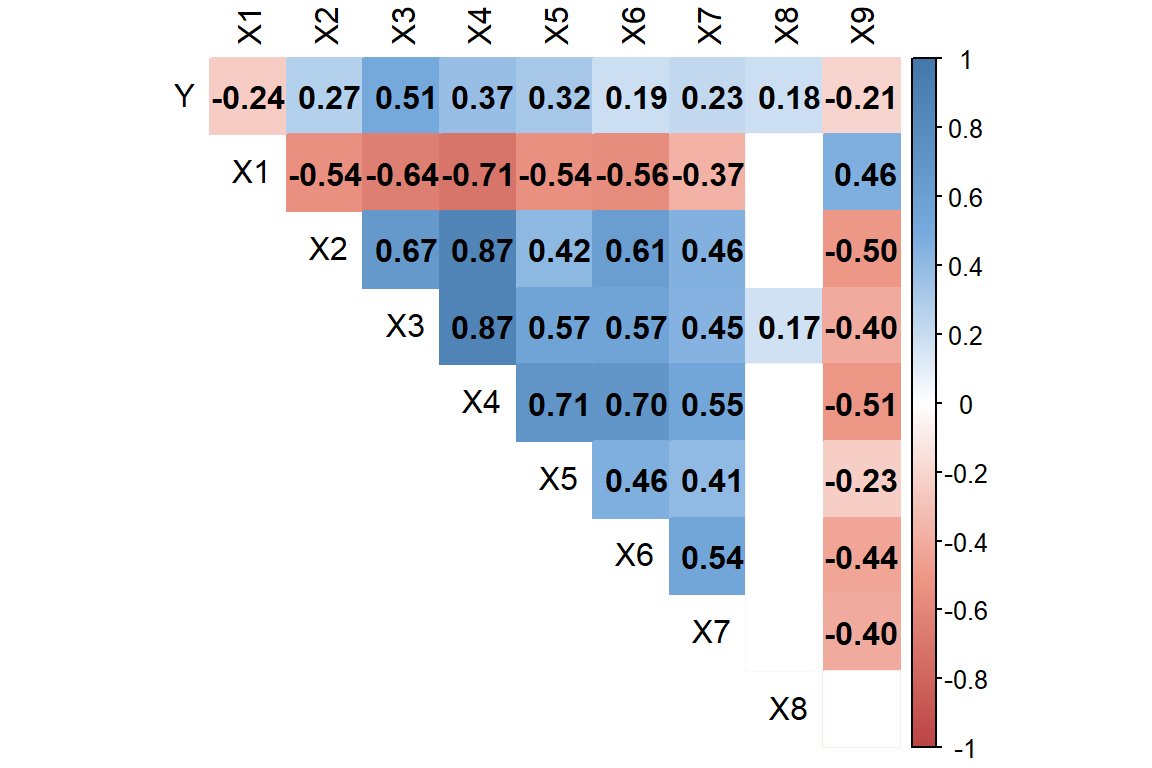
Peubah yang memiliki korelasi positif taraf sedang yaitu \(X_1\) dan \(X_5\) sedangkan \(X_2\) dan \(X_5\) memiliki korelasi negatif taraf sedang
Pada peubah \(X_2\) dengan \(X_4\), dan \(X_3\) dengan \(X_4\) terlihat korelasi yang cukup tinggi yaitu sebesar 0.87. Hal ini mengindikasikan adanya multikolinearitas.
Matriks Bobot
Matriks Pembobot berdasarkan jarak
pdrb1 <- pdrb
coordinates(pdrb1) <- ~longitude+latitude
plot(pdrb1)
#menghitung matriks jarak
longlat <- coordinates(pdrb1)
jarak<-as.matrix((dist(longlat)))K-Nearest Neighbour
# k=5, 5 tetangga terdekat
W.knn<-knn2nb(knearneigh(longlat,k=5,longlat=TRUE))
#> Warning in knn2nb(knearneigh(longlat, k = 5, longlat = TRUE)):
#> neighbour object has 3 sub-graphs
W.knn
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 2570
#> Percentage nonzero weights: 0.973
#> Average number of links: 5
#> 3 disjoint connected subgraphs
#> Non-symmetric neighbours listW.knn1 <- nb2listw(W.knn,style='W')
W.knn1
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 2570
#> Percentage nonzero weights: 0.973
#> Average number of links: 5
#> 3 disjoint connected subgraphs
#> Non-symmetric neighbours list
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 182 2126Inverse Distance Weight
#Alpha = 1
alpha1=1
W.idw <-1/(jarak^alpha1)
class(W.idw)
#> [1] "matrix" "array"#normalisasi matriks jarak
diag(W.idw)<-0
rtot<-rowSums(W.idw,na.rm=TRUE)
W.idw.sd<-W.idw/rtot #row-normalized
rowSums(W.idw.sd,na.rm=TRUE)
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 511 512 513 514
#> 1 1 1 1W.idw.1 = mat2listw(W.idw.sd,style='W')
summary(W.idw.1)
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 263682
#> Percentage nonzero weights: 99.8
#> Average number of links: 513
#> Link number distribution:
#>
#> 513
#> 514
#> 514 least connected regions:
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 with 513 links
#> 514 most connected regions:
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 with 513 links
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 20 2080#Alpha = 2
alpha2 = 2
W.idw2 <- 1/(jarak^alpha2)
#normalisasi baris
diag(W.idw2) <- 0
rtot <- rowSums(W.idw2,na.rm=TRUE)
W.idw.sd2 <- W.idw2/rtot #row-normalized
rowSums(W.idw.sd2,na.rm=TRUE)
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 511 512 513 514
#> 1 1 1 1
W.idw.22 = mat2listw(W.idw.sd2,style='W')
summary(W.idw.22)
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 263682
#> Percentage nonzero weights: 99.8
#> Average number of links: 513
#> Link number distribution:
#>
#> 513
#> 514
#> 514 least connected regions:
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 with 513 links
#> 514 most connected regions:
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 with 513 links
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 163 2123Eksponensial Distance Weight
alpha=1
W.exp <-exp((-alpha)*jarak)
diag(W.exp) <- 0
rtot<-rowSums(W.exp,na.rm=TRUE)
W.e.sd<-W.exp/rtot #row-normalized
rowSums(W.e.sd,na.rm=TRUE)
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510
#> 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1
#> 511 512 513 514
#> 1 1 1 1
W.ed1 = mat2listw(W.e.sd,style='W')
summary(W.ed1)
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 263682
#> Percentage nonzero weights: 99.8
#> Average number of links: 513
#> Link number distribution:
#>
#> 513
#> 514
#> 514 least connected regions:
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 with 513 links
#> 514 most connected regions:
#> 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 with 513 links
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 44 2072#peta
peta <- readOGR(dsn="data/shp/BATAS KABUPATEN KOTA DESEMBER 2019 DUKCAPIL.shp")
#> Warning in readOGR(dsn = "data/shp/BATAS KABUPATEN KOTA DESEMBER
#> 2019 DUKCAPIL.shp"): OGR support is provided by the sf and terra
#> packages among others
#> Warning in ogrListLayers(dsn = dsn): OGR support is provided by the
#> sf and terra packages among others
#> Warning in ogrInfo(dsn = dsn, layer = layer, encoding = encoding,
#> use_iconv = use_iconv, : OGR support is provided by the sf and
#> terra packages among others
#> Warning in ogrFIDs(dsn = dsn, layer = layer): OGR support is
#> provided by the sf and terra packages among others
#> Warning in OGRSpatialRef(dsn, layer, morphFromESRI = morphFromESRI,
#> dumpSRS = dumpSRS, : OGR support is provided by the sf and terra
#> packages among others
#> Warning in ogrListLayers(dsn): OGR support is provided by the sf
#> and terra packages among others
#> Warning in ogrFIDs(dsn = dsn, layer = layer): OGR support is
#> provided by the sf and terra packages among others
#> OGR data source with driver: ESRI Shapefile
#> Source: "C:\Users\anugraha\Documents\Materi_Orasi\book\data\shp\BATAS KABUPATEN KOTA DESEMBER 2019 DUKCAPIL.shp", layer: "BATAS KABUPATEN KOTA DESEMBER 2019 DUKCAPIL"
#> with 515 features
#> It has 1 fields
peta2 <- subset(peta, !is.na(peta@data$KAB_KOTA))
sf_use_s2(FALSE)
#> Spherical geometry (s2) switched offRook
Queen
Pemilihan Matriks Bobot
#knn
MI.knn <- moran(pdrb1$Y, W.knn1, n=length(W.knn1$neighbours), S0=Szero(W.knn1))
#radial
#MI.radial <- moran(pdrb1$Y, W.dmax.s, n=length(W.dmax.s$neighbours), S0=Szero(W.dmax.s))
#power distance alpha=1
MI.power1 <- moran(pdrb1$Y, W.idw.1, n=length(W.idw.1$neighbours), S0=Szero(W.idw.1))
#power distance alpha=2
MI.power2 <- moran(pdrb1$Y, W.idw.22, n=length(W.idw.22$neighbours), S0=Szero(W.idw.22))
#exponencial distance
MI.exp1 <- moran(pdrb1$Y, W.ed1, n=length(W.ed1$neighbours), S0=Szero(W.ed1))
#rook
MI.rook <- moran.test(pdrb1$Y, W.rook.s, randomisation = TRUE, zero.policy = TRUE)
MI.rook$estimate
#> Moran I statistic Expectation Variance
#> 0.01258 -0.00207 0.00107
#queen
MI.queen <- moran.test(pdrb1$Y, W.queen.s, randomisation = TRUE, zero.policy = TRUE)
MI.queen$estimate
#> Moran I statistic Expectation Variance
#> 0.01253 -0.00207 0.00107Perbandingan Nilai Moran Index
moranindeks<-data.frame(
"Matriks Bobot"=c("KNN (k=5)", "Power distance weight (alpha=1)", "Power distance weight (alpha=2)", "Exponential distance weight (alpha=1)", "Rook Contiguity", "Queen Contiguity"),
"Nilai Indeks Moran"=c(MI.knn$I, MI.power1$I, MI.power2$I,
MI.exp1$I, MI.rook[["estimate"]][["Moran I statistic"]] , MI.queen[["estimate"]][["Moran I statistic"]]))
moranindeks
#> Matriks.Bobot Nilai.Indeks.Moran
#> 1 KNN (k=5) 0.4407
#> 2 Power distance weight (alpha=1) 0.1837
#> 3 Power distance weight (alpha=2) 0.3530
#> 4 Exponential distance weight (alpha=1) 0.2262
#> 5 Rook Contiguity 0.0126
#> 6 Queen Contiguity 0.0125Woptimum <- W.knn1
moran.test(pdrb1$Y, Woptimum, randomisation = TRUE, zero.policy = TRUE)
#>
#> Moran I test under randomisation
#>
#> data: pdrb1$Y
#> weights: Woptimum
#>
#> Moran I statistic standard deviate = 18, p-value <2e-16
#> alternative hypothesis: greater
#> sample estimates:
#> Moran I statistic Expectation Variance
#> 0.440669 -0.001949 0.000625Model
ols = lm(Y~., data = datagwr)
summary(ols)
#>
#> Call:
#> lm(formula = Y ~ ., data = datagwr)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -1.04e+08 -1.68e+07 -2.01e+06 1.04e+07 3.39e+08
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 17223460 54103316 0.32 0.75036
#> X1 557589 341063 1.63 0.10270
#> X2 15983069 3276142 4.88 1.4e-06 ***
#> X3 19970 1728 11.56 < 2e-16 ***
#> X4 -10612244 1626242 -6.53 1.7e-10 ***
#> X5 6807112 1024833 6.64 8.0e-11 ***
#> X6 -188787 135979 -1.39 0.16564
#> X7 224613 137957 1.63 0.10412
#> X8 -175703 379097 -0.46 0.64322
#> X9 -1134771 338150 -3.36 0.00085 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 38600000 on 504 degrees of freedom
#> Multiple R-squared: 0.362, Adjusted R-squared: 0.351
#> F-statistic: 31.8 on 9 and 504 DF, p-value: <2e-16Uji Asumsi Model
Normality Test
Plot sisaan model Regresi Klasik
sisaan.ols <-residuals(ols)
hist(sisaan.ols,
xlab = "sisaan",
col = "#27D3D3",
breaks=30,
prob = TRUE)
lines(density(sisaan.ols), # density plot
lwd = 2, # thickness of line
col = "chocolate3")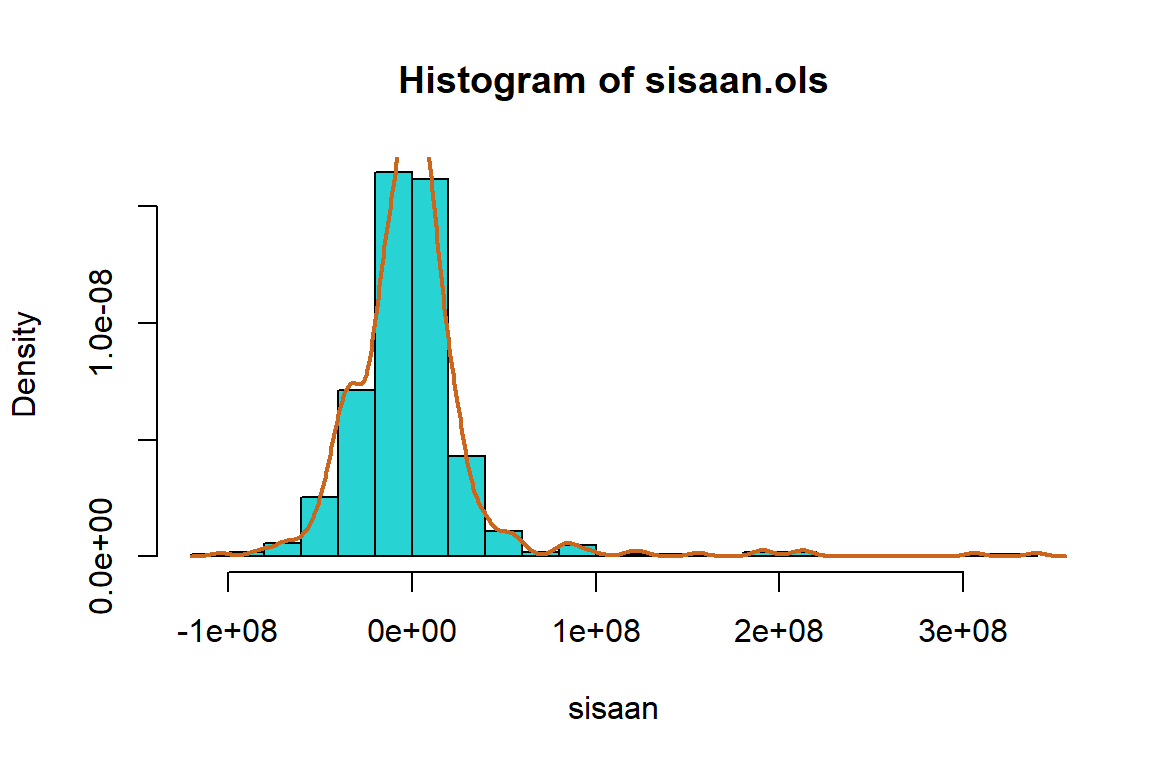
qqnorm(sisaan.ols,datax=T, col="blue")
qqline(rnorm(length(sisaan.ols),mean(sisaan.ols),sd(sisaan.ols)),datax=T, col="red")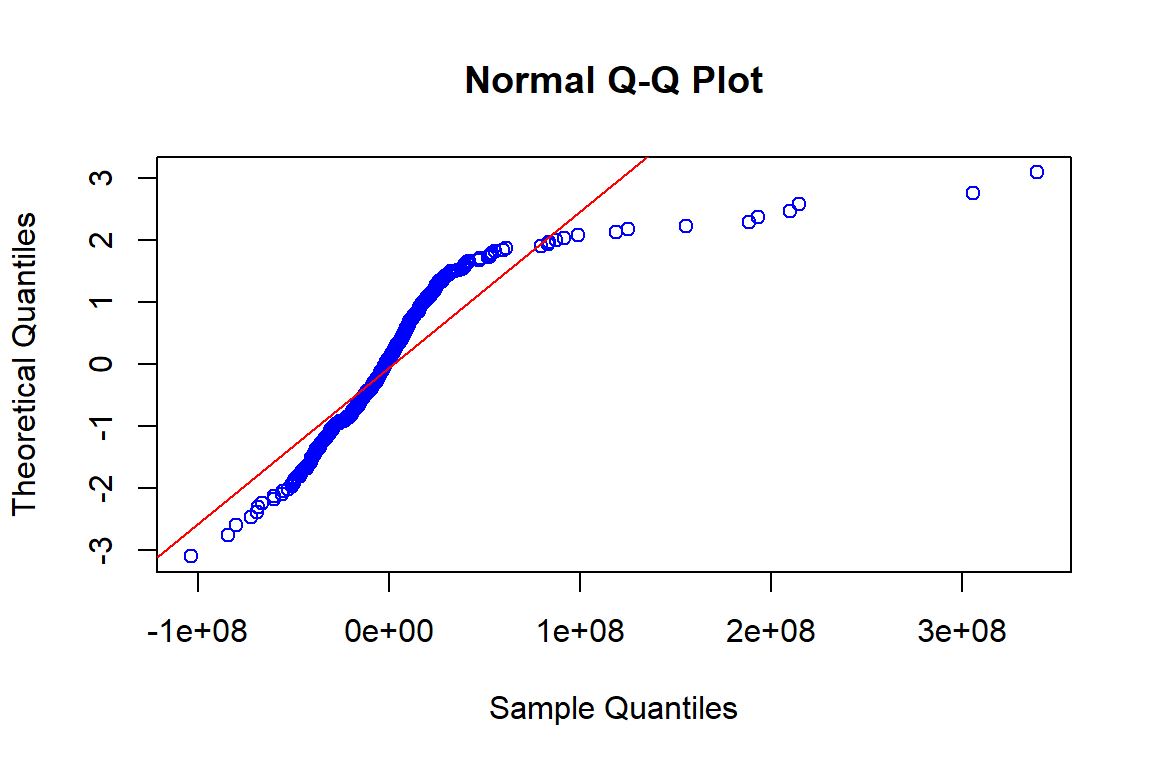
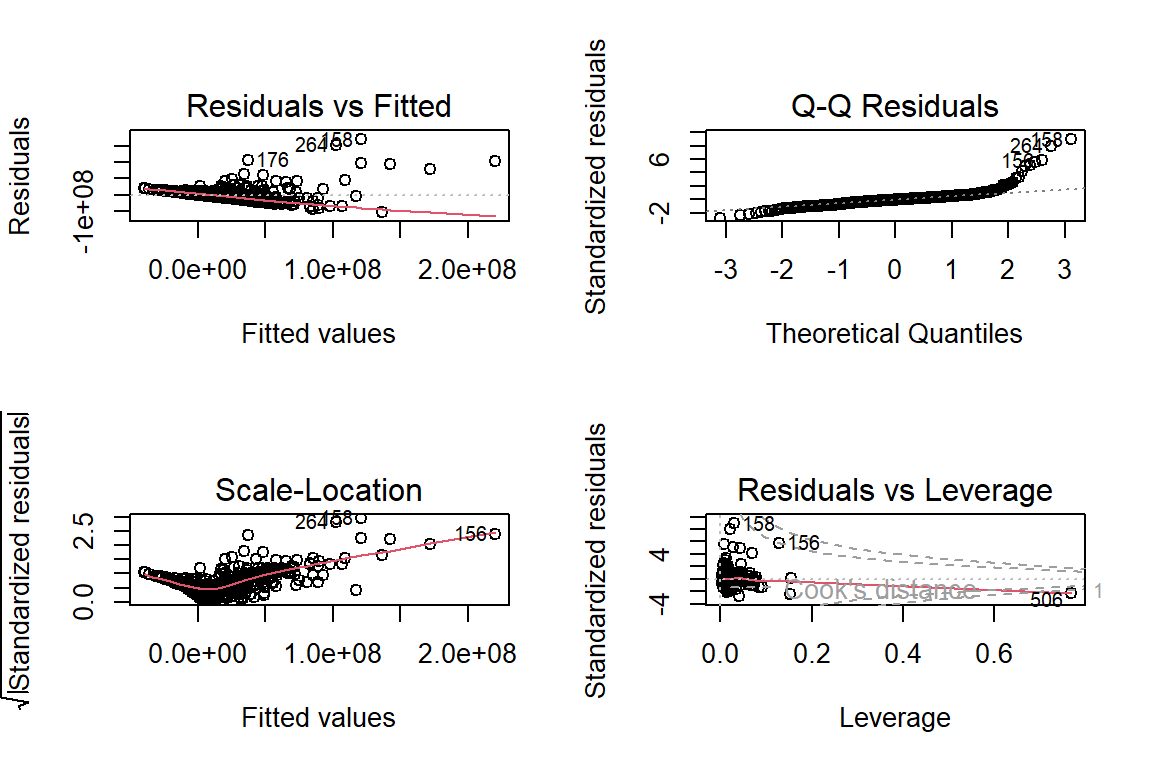
Berdasarkan plot histogram tampak bahwa data cenderung menjulur ke kanan dan pada plot QQ normal cukup banyak titik-titik yang tidak berada di sekitar garis yang mengindikasikan secara eksploratif asumsi normalitas belum terpenuhi.
ad.test(sisaan.ols)
#>
#> Anderson-Darling normality test
#>
#> data: sisaan.ols
#> A = 26, p-value <2e-16
jarque.bera.test(sisaan.ols)
#>
#> Jarque Bera Test
#>
#> data: sisaan.ols
#> X-squared = 14576, df = 2, p-value <2e-16Hipotesis yang digunakan adalah
\(H_0\): sisaan model menyebar normal \(H_1\): sisaan model tidak menyebar normal
Hasil dari pengujian asumsi normlitas nilai p value yang diperoleh < alpha 5% baik dengan jarque bera ataupun anderson darling. Sehingga tolak \(H_0\) yang menandakan bahwa sisaan belum menyebar normal. (Asumsi kenormalam belum terpenuhi)
Heteroscedasticity
bptest(ols)
#>
#> studentized Breusch-Pagan test
#>
#> data: ols
#> BP = 103, df = 9, p-value <2e-16Hipotesis yang digunakan adalah
\(H_0\): ragam sisaan homogen \(H_1\): ragam sisaan tidak homogen
Nilai p-value yang diperoleh < alpha (5%) sehingga tolak \(H_0\) dan dapat disimpulkan bahwa ragam dari sisaan tidak homogen.
VIF
vif(ols)
#> X1 X2 X3 X4 X5 X6 X7 X8 X9
#> 2.23 9.83 7.59 38.45 4.30 2.20 1.61 1.23 1.61Berdasarkan pengujian vif untuk melihat adanya multikolinearitas, diperoleh nilai vif yang > 5 pada peubah \(X_2\), \(X_3\), \(X_4\) yang mengindikasikan terdapat multikolinearitas.
Autocorrelation
Dengan durbin watson, tanpa memasukkan weight
dwtest(ols)
#>
#> Durbin-Watson test
#>
#> data: ols
#> DW = 1, p-value <2e-16
#> alternative hypothesis: true autocorrelation is greater than 0Hipotesis yang digunakan adalah
\(H_0\): tidak ada autokorelasi \(H_1\): terdapat autokorelasi
Nilai p=value yang diperoleh 2.2e-16 < alpha (5%) sehingga tolak \(H_0\) sehingga dapat dikatakan terdapat gejala autokorelasi pada sisaan.
Melihat autokorelasi dengan moran test dan matriks jarak yang digunakan adalah KNN dengan \(k=5\).
lm.morantest(ols, W.knn1 , alternative="two.sided")
#>
#> Global Moran I for regression residuals
#>
#> data:
#> model: lm(formula = Y ~ ., data = datagwr)
#> weights: W.knn1
#>
#> Moran I statistic standard deviate = 11, p-value <2e-16
#> alternative hypothesis: two.sided
#> sample estimates:
#> Observed Moran I Expectation Variance
#> 0.260992 -0.008560 0.000651moran.test(sisaan.ols, W.knn1 ,randomisation=F, alternative="two.sided")
#>
#> Moran I test under normality
#>
#> data: sisaan.ols
#> weights: W.knn1
#>
#> Moran I statistic standard deviate = 10, p-value <2e-16
#> alternative hypothesis: two.sided
#> sample estimates:
#> Moran I statistic Expectation Variance
#> 0.260992 -0.001949 0.000679Hipotesis yang digunakan :
\(H_0\) : Tidak terdapat autokorelasi pada sisaan \(H_1\) : Terdapat autokorelasi pada sisaan
Berdasarkan output diatas, hasil pengujian dengan lm.moran test dan moran.test diperoleh p-value yang sangat kecil. Nilai p-value < alpha (5%) sehingga dapat menolak \(H_0\) Artinya kita dapat menyimpulkan bahwa terdapat cukup bukti untuk menyatakan bahwa terdapat autokorelasi pada sisaan model OLS dengan taraf nyata 5%. Atau dengan kata lain, model dengan regresi klasik saja tidak cukup, karena terjadi dependensi spasial sehingga diperlukan uji Lagrange Multiplier (LM) untuk mengidentifikasi model regresi dependensi spasial yang sesuai.
Seleksi peubah
Model
datgwr2 <- datagwr %>%
dplyr::select(-c('X2','X3','X4'))
ols1 = lm(log(Y)~., data = datgwr2)
summary(ols1)
#>
#> Call:
#> lm(formula = log(Y) ~ ., data = datgwr2)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.2482 -0.6427 -0.0234 0.5300 2.8270
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 7.50829 1.13690 6.60 1.0e-10 ***
#> X1 -0.03650 0.00756 -4.83 1.8e-06 ***
#> X5 0.14510 0.01497 9.69 < 2e-16 ***
#> X6 -0.00291 0.00304 -0.96 0.33959
#> X7 0.01064 0.00327 3.25 0.00124 **
#> X8 0.03958 0.00832 4.76 2.5e-06 ***
#> X9 -0.02947 0.00773 -3.81 0.00015 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 0.93 on 507 degrees of freedom
#> Multiple R-squared: 0.469, Adjusted R-squared: 0.462
#> F-statistic: 74.5 on 6 and 507 DF, p-value: <2e-16Uji Asumsi
Normality Test
Plot sisaan model Regresi Klasik
sisaan.ols1 <-residuals(ols1)
hist(sisaan.ols1,
xlab = "sisaan",
col = "#27D3D3",
breaks=30,
prob = TRUE)
lines(density(sisaan.ols1), # density plot
lwd = 2, # thickness of line
col = "chocolate3")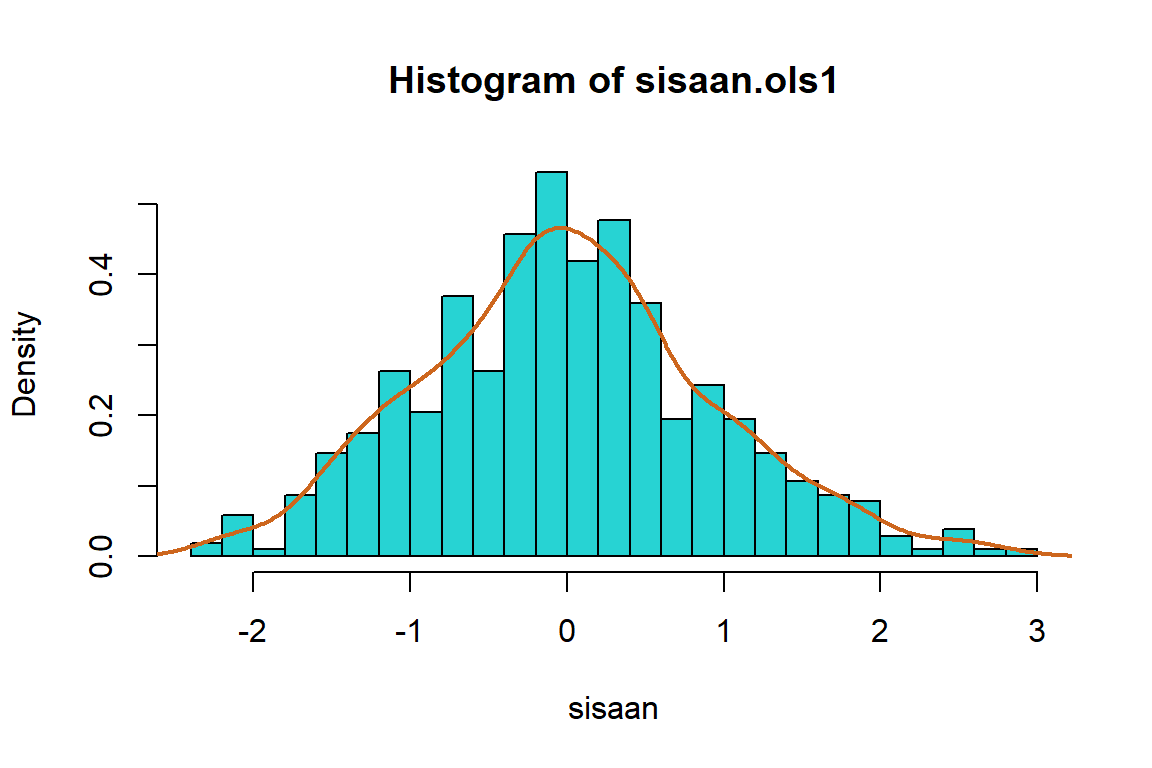
qqnorm(sisaan.ols1,datax=T, col="blue")
qqline(rnorm(length(sisaan.ols1),mean(sisaan.ols1),sd(sisaan.ols1)),datax=T, col="red")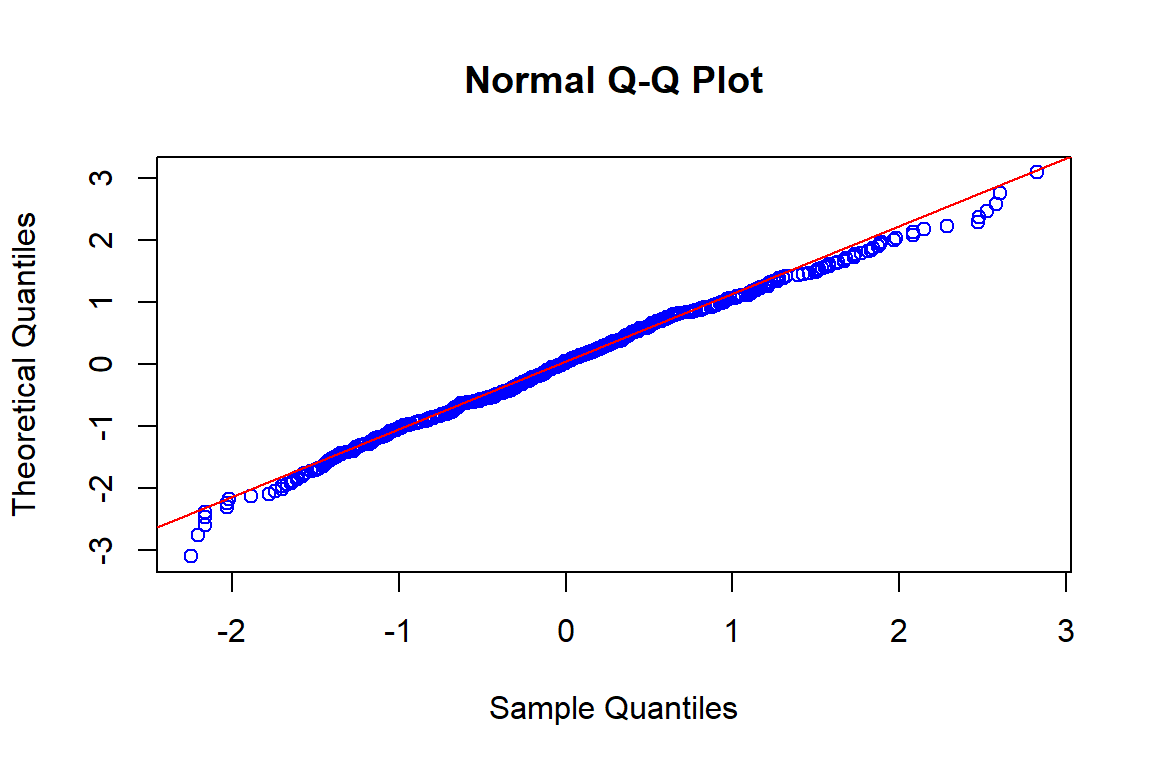
Berdasarkan plot histogram tampak bahwa data sudah membentuk pola simetris dan pada plot QQ normal cukup banyak titik-titik yang berada di sekitar garis lurus yang mengindikasikan secara eksploratif asumsi normalitas terpenuhi.
ad.test(sisaan.ols1)
#>
#> Anderson-Darling normality test
#>
#> data: sisaan.ols1
#> A = 0.6, p-value = 0.09
jarque.bera.test(sisaan.ols1)
#>
#> Jarque Bera Test
#>
#> data: sisaan.ols1
#> X-squared = 4, df = 2, p-value = 0.1Hipotesis yang digunakan adalah
\(H_0\): sisaan model menyebar normal \(H_1\): sisaan model tidak menyebar normal
Hasil dari pengujian asumsi normalitas nilai p value yang diperoleh > alpha 5% anderson darling. Sehingga terima \(H_0\) yang menandakan bahwa sisaanmenyebar normal. (Asumsi kenormalan terpenuhi)
Heteroscedasticity
bptest(ols1)
#>
#> studentized Breusch-Pagan test
#>
#> data: ols1
#> BP = 29, df = 6, p-value = 5e-05Hipotesis yang digunakan adalah
\(H_0\): ragam sisaan homogen \(H_1\): ragam sisaan tidak homogen
Nilai p-value yang diperoleh < alpha (5%) sehingga tolak \(H_0\) dan dapat disimpulkan bahwa ragam dari sisaan tidak homogen.
Autocorrelation
Dengan Durbin Watson, tanpa memasukkan bobot
dwtest(ols1)
#>
#> Durbin-Watson test
#>
#> data: ols1
#> DW = 1, p-value <2e-16
#> alternative hypothesis: true autocorrelation is greater than 0Hipotesis yang digunakan adalah
\(H_0\): tidak ada autokorelasi \(H_1\): terdapat autokorelasi
Nilai p-value yang diperoleh 2.2e-16, < alpha (5%) sehingga tolak \(H_0\) sehingga dapat dikatakan terdapat gejala autokorelasi pada sisaan.
vif(ols1)
#> X1 X5 X6 X7 X8 X9
#> 1.88 1.58 1.89 1.56 1.02 1.45Setelah dilakukan reduksi peubah \(X_2\), \(X_3\), dan \(X_4\) tidak ditemukan adanya masalah multikolinearitas. Setelah dilakukan treatment berupa log pada Y dan membuang peubah \(X_2\), \(X_3\), dan \(X_4\), asumsi kenormalan dan VIF dapat tertangani.Akan tetapi untuk asumsi kebebasan sisaan dan kehomogenan ragam belum dapat tertangani. Oleh karena itu diperlukan pemodelan lain yang dapat mengakomodir hal tersebut yaitu Mixed GWR.
UJI LM
Pengujian dilakukan untuk melihat k optimum yang dapat digunakan untuk memberikan model terbaik.
Menentukan k atau NN optimal.
#menghitung matriks jarak
longlat <- coordinates(pdrb1)
jarak<-as.matrix((dist(longlat)))#k=1, 1 tetangga terdekat
W.knn1<-knn2nb(knearneigh(longlat,k=1,longlat=TRUE)) #matriks bobot dengan knn k=1 #knearneigh(x, k=1, longlat = NULL, use_kd_tree=TRUE)
#> Warning in knn2nb(knearneigh(longlat, k = 1, longlat = TRUE)):
#> neighbour object has 147 sub-graphs
W.knn1
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 514
#> Percentage nonzero weights: 0.195
#> Average number of links: 1
#> 147 disjoint connected subgraphs
#> Non-symmetric neighbours list
W.knn1a <- nb2listw(W.knn1,style='W')
W.knn1a
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 514
#> Percentage nonzero weights: 0.195
#> Average number of links: 1
#> 147 disjoint connected subgraphs
#> Non-symmetric neighbours list
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 808 2418
# LM-test
LM1<-lm.RStests(ols1, W.knn1a, test=c("RSlag"))
summary(LM1)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: W.knn1a
#>
#> statistic parameter p.value
#> RSlag 38.9 1 4.4e-10 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=2,
W.knn2<-knn2nb(knearneigh(longlat,k=2,longlat=TRUE))
#> Warning in knn2nb(knearneigh(longlat, k = 2, longlat = TRUE)):
#> neighbour object has 27 sub-graphs
W.knn2
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 1028
#> Percentage nonzero weights: 0.389
#> Average number of links: 2
#> 27 disjoint connected subgraphs
#> Non-symmetric neighbours list
W.knn2a <- nb2listw(W.knn2,style='W')
W.knn2a
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 1028
#> Percentage nonzero weights: 0.389
#> Average number of links: 2
#> 27 disjoint connected subgraphs
#> Non-symmetric neighbours list
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 434 2210
# LM-test
LM2<-lm.RStests(ols1, W.knn2a,
test=c("RSlag"))
summary(LM2)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: W.knn2a
#>
#> statistic parameter p.value
#> RSlag 60.9 1 5.9e-15 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=3,
W.knn3<-knn2nb(knearneigh(longlat,k=3,longlat=TRUE))
#> Warning in knn2nb(knearneigh(longlat, k = 3, longlat = TRUE)):
#> neighbour object has 8 sub-graphs
W.knn3
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 1542
#> Percentage nonzero weights: 0.584
#> Average number of links: 3
#> 8 disjoint connected subgraphs
#> Non-symmetric neighbours list
W.knn3a <- nb2listw(W.knn3,style='W')
W.knn3a
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 1542
#> Percentage nonzero weights: 0.584
#> Average number of links: 3
#> 8 disjoint connected subgraphs
#> Non-symmetric neighbours list
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 298 2156
# LM-test
LM3<-lm.LMtests(ols1, W.knn3a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM3)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 83.3 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=4,
W.knn4<-knn2nb(knearneigh(longlat,k=4,longlat=TRUE))
#> Warning in knn2nb(knearneigh(longlat, k = 4, longlat = TRUE)):
#> neighbour object has 4 sub-graphs
W.knn4
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 2056
#> Percentage nonzero weights: 0.778
#> Average number of links: 4
#> 4 disjoint connected subgraphs
#> Non-symmetric neighbours list
W.knn4a <- nb2listw(W.knn4,style='W')
W.knn4a
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 2056
#> Percentage nonzero weights: 0.778
#> Average number of links: 4
#> 4 disjoint connected subgraphs
#> Non-symmetric neighbours list
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 224 2138
# LM-test
LM4<-lm.LMtests(ols1, W.knn4a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM4)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 109 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=5,
W.knn5<-knn2nb(knearneigh(longlat,k=5,longlat=TRUE))
#> Warning in knn2nb(knearneigh(longlat, k = 5, longlat = TRUE)):
#> neighbour object has 3 sub-graphs
W.knn5
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 2570
#> Percentage nonzero weights: 0.973
#> Average number of links: 5
#> 3 disjoint connected subgraphs
#> Non-symmetric neighbours list
W.knn5a <- nb2listw(W.knn5,style='W')
W.knn5a
#> Characteristics of weights list object:
#> Neighbour list object:
#> Number of regions: 514
#> Number of nonzero links: 2570
#> Percentage nonzero weights: 0.973
#> Average number of links: 5
#> 3 disjoint connected subgraphs
#> Non-symmetric neighbours list
#>
#> Weights style: W
#> Weights constants summary:
#> n nn S0 S1 S2
#> W 514 264196 514 182 2126
# LM-test
LM5<-lm.LMtests(ols1, W.knn5a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM5)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 118 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=6,
W.knn6<-knn2nb(knearneigh(longlat,k=6,longlat=TRUE))
#> Warning in knn2nb(knearneigh(longlat, k = 6, longlat = TRUE)):
#> neighbour object has 2 sub-graphs
W.knn6a <- nb2listw(W.knn6,style='W')
# LM-test
LM6<-lm.LMtests(ols1, W.knn6a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM6)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 139 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=7,
W.knn7<-knn2nb(knearneigh(longlat,k=7,longlat=TRUE))
#> Warning in knn2nb(knearneigh(longlat, k = 7, longlat = TRUE)):
#> neighbour object has 2 sub-graphs
W.knn7a <- nb2listw(W.knn7,style='W')
# LM-test
LM7<-lm.LMtests(ols1, W.knn7a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM7)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 142 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=8,
W.knn8<-knn2nb(knearneigh(longlat,k=8,longlat=TRUE))
W.knn8a <- nb2listw(W.knn8,style='W')
# LM-test
LM8<-lm.LMtests(ols1, W.knn8a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM8)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 149 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=9,
W.knn9<-knn2nb(knearneigh(longlat,k=9,longlat=TRUE))
W.knn9a <- nb2listw(W.knn8,style='W')
# LM-test
LM9<-lm.LMtests(ols1, W.knn9a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM9)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 149 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1#k=10,
W.knn10<-knn2nb(knearneigh(longlat,k=10,longlat=TRUE))
W.knn10a <- nb2listw(W.knn10,style='W')
# LM-test
LM10<-lm.LMtests(ols1, W.knn10a,
test=c("RSlag"))
#> Please update scripts to use lm.RStests in place of lm.LMtests
summary(LM10)
#> Rao's score (a.k.a Lagrange multiplier) diagnostics for
#> spatial dependence
#> data:
#> model: lm(formula = log(Y) ~ ., data = datgwr2)
#> test weights: listw
#>
#> statistic parameter p.value
#> RSlag 171 1 <2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Rangkuman k atau NN optimum
LM_k <- data.frame(k = c(1,2,3,4,5,6,7,8,9,10),
LMstat= c(round(LM1$RSlag$statistic,3),
round(LM2$RSlag$statistic,3),
round(LM3$RSlag$statistic,3),
round(LM4$RSlag$statistic,3),
round(LM5$RSlag$statistic,3),
round(LM6$RSlag$statistic,3),
round(LM7$RSlag$statistic,3),
round(LM8$RSlag$statistic,3),
round(LM9$RSlag$statistic,3),
round(LM10$RSlag$statistic,3)),
pvalue = c((LM1$RSlag$p.value),
(LM2$RSlag$p.value),
(LM3$RSlag$p.value),
(LM4$RSlag$p.value),
(LM5$RSlag$p.value),
(LM6$RSlag$p.value),
(LM7$RSlag$p.value),
(LM8$RSlag$p.value),
(LM9$RSlag$p.value),
(LM10$RSlag$p.value)
)
)
LM_k
#> k LMstat pvalue
#> 1 1 38.9 4.36e-10
#> 2 2 60.9 5.88e-15
#> 3 3 83.3 0.00e+00
#> 4 4 109.2 0.00e+00
#> 5 5 118.3 0.00e+00
#> 6 6 139.4 0.00e+00
#> 7 7 142.0 0.00e+00
#> 8 8 149.1 0.00e+00
#> 9 9 149.1 0.00e+00
#> 10 10 170.6 0.00e+00LM_k %>%
ggplot(aes(k, LMstat))+
ylim(0,180)+
geom_line(color="blue") +
geom_point(shape = 21, colour = "darkblue", fill = "red", size=2)+
geom_point(data = LM_k %>% filter(LMstat != max(LMstat)),
pch = 21, fill = "white", colour = "black")+
ggtitle("Nilai LMStat pada k = 1 s.d 10")+
theme_bw() +
annotate("text", x=9.5, y=175, label= " Opt(k=10)") 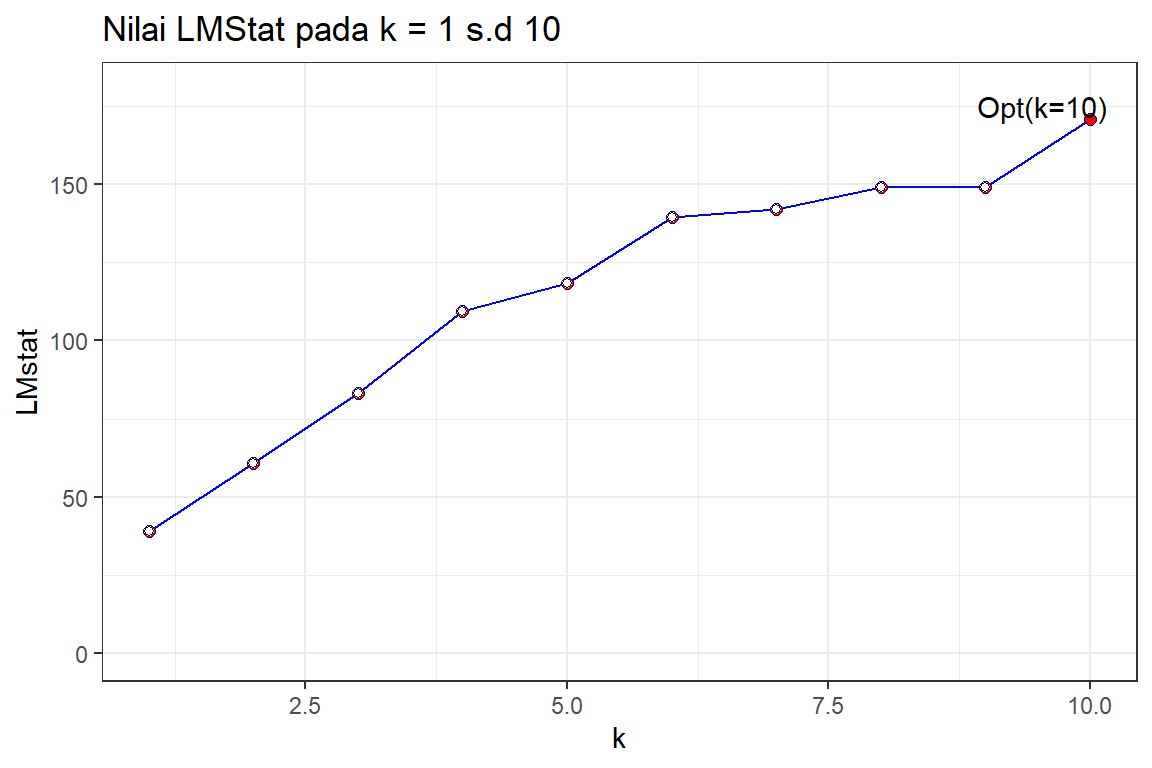
Berdasarkan pengujian dengan LMTest pada model SAR dan bobot KNN dengan k=2 sd. 10 diperoleh K = 10 yang memiliki LMstat terbesar yaitu 161.692.
GWR
Bandwith
Bisquare
#bw_bisquare
bwbs1<- gwr.sel(log(Y) ~.,data=datgwr2,
coords=cbind(pdrb$latitude,pdrb$longitude),gweight=gwr.bisquare, adapt=FALSE)
#> Bandwidth: 18.5 CV score: 395
#> Bandwidth: 29.9 CV score: 424
#> Bandwidth: 11.5 CV score: 375
#> Bandwidth: 7.1 CV score: 368
#> Bandwidth: 4.41 CV score: NA
#> Bandwidth: 8.77 CV score: 364
#> Bandwidth: 8.68 CV score: 364
#> Bandwidth: 8.38 CV score: 364
#> Bandwidth: 7.89 CV score: 364
#> Bandwidth: 8.31 CV score: 364
#> Bandwidth: 8.3 CV score: 364
#> Bandwidth: 8.3 CV score: 364
#> Bandwidth: 8.3 CV score: 364
#> Bandwidth: 8.3 CV score: 364
#> Bandwidth: 8.3 CV score: 364
#> Bandwidth: 8.3 CV score: 364modelgwr_bisq <- gwr(log(Y) ~.,data=datgwr2,
coords=cbind(pdrb$longitude,pdrb$latitude),
gweight=gwr.bisquare,
hatmatrix=TRUE,
bandwidth=bwbs1,
se.fit=TRUE)
#cek bw
bw <- modelgwr_bisq$bandwidth
bw
#> [1] 8.3
modelgwr_bisq
#> Call:
#> gwr(formula = log(Y) ~ ., data = datgwr2, coords = cbind(pdrb$longitude,
#> pdrb$latitude), bandwidth = bwbs1, gweight = gwr.bisquare,
#> hatmatrix = TRUE, se.fit = TRUE)
#> Kernel function: gwr.bisquare
#> Fixed bandwidth: 8.3
#> Summary of GWR coefficient estimates at data points:
#> Min. 1st Qu. Median 3rd Qu. Max.
#> X.Intercept. 6.44e+00 1.04e+01 1.19e+01 1.33e+01 1.63e+01
#> X1 -1.01e-01 -4.01e-02 -1.73e-02 3.38e-03 6.19e-02
#> X5 8.63e-03 5.29e-02 6.65e-02 7.95e-02 1.29e-01
#> X6 -1.68e-02 -6.03e-05 1.70e-03 5.48e-03 3.08e-02
#> X7 -2.57e-02 3.43e-03 1.24e-02 1.49e-02 2.52e-02
#> X8 -9.93e-03 3.69e-02 1.28e-01 1.47e-01 2.18e-01
#> X9 -7.54e-02 -3.44e-02 -2.85e-02 -1.34e-02 4.85e-02
#> Global
#> X.Intercept. 7.51
#> X1 -0.04
#> X5 0.15
#> X6 0.00
#> X7 0.01
#> X8 0.04
#> X9 -0.03
#> Number of data points: 514
#> Effective number of parameters (residual: 2traceS - traceS'S): 69.4
#> Effective degrees of freedom (residual: 2traceS - traceS'S): 445
#> Sigma (residual: 2traceS - traceS'S): 0.813
#> Effective number of parameters (model: traceS): 54.9
#> Effective degrees of freedom (model: traceS): 459
#> Sigma (model: traceS): 0.8
#> Sigma (ML): 0.756
#> AICc (GWR p. 61, eq 2.33; p. 96, eq. 4.21): 1297
#> AIC (GWR p. 96, eq. 4.22): 1226
#> Residual sum of squares: 294
#> Quasi-global R2: 0.644Gauss
bwgauss <- gwr.sel(log(Y) ~.,data=datgwr2,
coords=cbind(pdrb$longitude,pdrb$latitude),gweight=gwr.Gauss, adapt = F)
#> Bandwidth: 18.5 CV score: 449
#> Bandwidth: 29.9 CV score: 472
#> Bandwidth: 11.5 CV score: 417
#> Bandwidth: 7.1 CV score: 391
#> Bandwidth: 4.41 CV score: 370
#> Bandwidth: 2.74 CV score: 359
#> Bandwidth: 1.71 CV score: 381
#> Bandwidth: 3.25 CV score: 360
#> Bandwidth: 2.61 CV score: 359
#> Bandwidth: 2.85 CV score: 359
#> Bandwidth: 2.82 CV score: 359
#> Bandwidth: 2.82 CV score: 359
#> Bandwidth: 2.82 CV score: 359
#> Bandwidth: 2.82 CV score: 359
#> Bandwidth: 2.82 CV score: 359
#> Bandwidth: 2.82 CV score: 359
bwgauss
#> [1] 2.82modelgwr_gauss <- gwr(log(Y)~.,data=datgwr2,
bandwidth=bwgauss, coords=cbind(pdrb$longitude,pdrb$latitude),
hatmatrix=TRUE,gweight=gwr.Gauss,
se.fit = TRUE)
modelgwr_gauss
#> Call:
#> gwr(formula = log(Y) ~ ., data = datgwr2, coords = cbind(pdrb$longitude,
#> pdrb$latitude), bandwidth = bwgauss, gweight = gwr.Gauss,
#> hatmatrix = TRUE, se.fit = TRUE)
#> Kernel function: gwr.Gauss
#> Fixed bandwidth: 2.82
#> Summary of GWR coefficient estimates at data points:
#> Min. 1st Qu. Median 3rd Qu. Max.
#> X.Intercept. 5.370177 10.391929 12.047338 13.315224 17.710820
#> X1 -0.097845 -0.042252 -0.018471 0.005261 0.061538
#> X5 0.001216 0.050011 0.065041 0.083207 0.156672
#> X6 -0.012555 -0.000905 0.001939 0.006029 0.030969
#> X7 -0.024062 0.003228 0.012895 0.014368 0.023899
#> X8 -0.021298 0.041204 0.119960 0.154086 0.222766
#> X9 -0.085776 -0.035204 -0.029201 -0.013140 0.046274
#> Global
#> X.Intercept. 7.51
#> X1 -0.04
#> X5 0.15
#> X6 0.00
#> X7 0.01
#> X8 0.04
#> X9 -0.03
#> Number of data points: 514
#> Effective number of parameters (residual: 2traceS - traceS'S): 91.8
#> Effective degrees of freedom (residual: 2traceS - traceS'S): 422
#> Sigma (residual: 2traceS - traceS'S): 0.803
#> Effective number of parameters (model: traceS): 69.3
#> Effective degrees of freedom (model: traceS): 445
#> Sigma (model: traceS): 0.782
#> Sigma (ML): 0.728
#> AICc (GWR p. 61, eq 2.33; p. 96, eq. 4.21): 1295
#> AIC (GWR p. 96, eq. 4.22): 1201
#> Residual sum of squares: 272
#> Quasi-global R2: 0.67Rangkuman Model GWR
Bandwith dengan gauss memberikan nilai AIC yang sedikit lebih baik. Sehingga bandwith optimal dengan gauss (bwgauss) akan digunakan untuk membentuk matriks pembobot W_gauss.
MGWR
#Menyiapkan koordinat
coord <- as.matrix(pdrb[,c("latitude","longitude")])
#menyiapkan
## Creating a spatial weight matrix (sparce dgCMatrix) of 10 nearest neighbors with 0 in diagonal
# H = bandwith optimal dengan gauss, NN=10 merupakan k optimal dari hasil LM-test dengan KNN
W_gauss <- kernel_matW(
H = bwgauss,
kernels = 'gauss',
coord_i = coord,
NN = 10,
adaptive = TRUE,
diagnull = TRUE,
rowNorm = TRUE
)
# H = bandwith optimal dengan bisquare, NN=10 merupakan k optimal dari hasil LM-test dengan KNN
W_bisq <- kernel_matW(
H = bwbs1,
kernels = 'bisq',
coord_i = coord,
NN = 10,
adaptive = TRUE,
diagnull = TRUE,
rowNorm = TRUE
)Model 1 OLS
summary(ols1)
#>
#> Call:
#> lm(formula = log(Y) ~ ., data = datgwr2)
#>
#> Residuals:
#> Min 1Q Median 3Q Max
#> -2.2482 -0.6427 -0.0234 0.5300 2.8270
#>
#> Coefficients:
#> Estimate Std. Error t value Pr(>|t|)
#> (Intercept) 7.50829 1.13690 6.60 1.0e-10 ***
#> X1 -0.03650 0.00756 -4.83 1.8e-06 ***
#> X5 0.14510 0.01497 9.69 < 2e-16 ***
#> X6 -0.00291 0.00304 -0.96 0.33959
#> X7 0.01064 0.00327 3.25 0.00124 **
#> X8 0.03958 0.00832 4.76 2.5e-06 ***
#> X9 -0.02947 0.00773 -3.81 0.00015 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> Residual standard error: 0.93 on 507 degrees of freedom
#> Multiple R-squared: 0.469, Adjusted R-squared: 0.462
#> F-statistic: 74.5 on 6 and 507 DF, p-value: <2e-16Model 2 GWR
opt.bandwith.gwr <- bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord=coord,
fixed_vars='Intercept',
Models='GWR',
candidates_Kernels=c('bisq','gauss'),
control=list(),
control_search=list()
)H_bisq <- opt.bandwith.gwr[["GWR_bisq_adaptive"]][["config_model"]][["H"]]
H_gauss <- opt.bandwith.gwr[["GWR_gauss_adaptive"]][["config_model"]][["H"]]gwr_bisq <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
kernels = c('bisq'),
H = H_bisq,
Model = 'GWR',
control = list(SE=TRUE,adaptive=TRUE)
)
summary_mgwrsar(gwr_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> kernels = c("bisq"), H = H_bisq, Model = "GWR", control = list(SE = TRUE,
#> adaptive = TRUE))
#> Model: GWR
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 131
#> Computation time: 0.62
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Varying parameters: Intercept X1 X5 X6 X7 X8 X9
#> Intercept X1 X5 X6 X7 X8
#> Min. 4.41515 -0.08698 -0.00463 -0.02221 -0.01422 -0.01728
#> 1st Qu. 10.38515 -0.04664 0.03921 -0.00119 0.00456 0.02427
#> Median 12.66404 -0.01862 0.06172 0.00330 0.00929 0.10316
#> Mean 12.37147 -0.02074 0.06228 0.00442 0.00730 0.09803
#> 3rd Qu. 14.60060 0.00865 0.08255 0.00725 0.01224 0.14648
#> Max. 18.34132 0.05908 0.17277 0.03146 0.02839 0.22430
#> X9
#> Min. -0.05
#> 1st Qu. -0.04
#> Median -0.03
#> Mean -0.03
#> 3rd Qu. -0.02
#> Max. 0.04
#> Effective degrees of freedom: 454
#> AICc: 1447
#> Residual sum of squares: 287
#> RMSE: 0.748gwr_gauss <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
kernels = c('gauss'),
H = H_gauss,
Model = 'GWR',
control = list(SE=TRUE,adaptive=TRUE)
)
summary_mgwrsar(gwr_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> kernels = c("gauss"), H = H_gauss, Model = "GWR", control = list(SE = TRUE,
#> adaptive = TRUE))
#> Model: GWR
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 48
#> Computation time: 0.72
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Varying parameters: Intercept X1 X5 X6 X7 X8 X9
#> Intercept X1 X5 X6 X7 X8
#> Min. 5.63e+00 -9.46e-02 6.67e-03 -8.24e-03 -8.72e-03 -2.31e-02
#> 1st Qu. 8.99e+00 -3.59e-02 5.23e-02 -2.55e-05 4.39e-03 2.82e-02
#> Median 1.12e+01 -1.93e-02 7.86e-02 2.48e-03 9.64e-03 1.01e-01
#> Mean 1.14e+01 -2.50e-02 7.65e-02 4.00e-03 8.05e-03 9.32e-02
#> 3rd Qu. 1.40e+01 -4.43e-03 1.02e-01 5.88e-03 1.31e-02 1.46e-01
#> Max. 1.81e+01 2.76e-02 1.48e-01 2.69e-02 1.84e-02 1.88e-01
#> X9
#> Min. -0.05
#> 1st Qu. -0.04
#> Median -0.03
#> Mean -0.03
#> 3rd Qu. -0.01
#> Max. 0.00
#> Effective degrees of freedom: 468
#> AICc: 1427
#> Residual sum of squares: 303
#> RMSE: 0.768Model_GWR <- c("Bisquare Adaptive", "Gauss Adaptive")
AIC_GWR <- c(1219.306, 1233.073)
RMSE_GWR <- c(gwr_bisq$RMSE, gwr_gauss$RMSE)
data.frame(Model_GWR, AIC_GWR, RMSE_GWR)
#> Model_GWR AIC_GWR RMSE_GWR
#> 1 Bisquare Adaptive 1219 0.748
#> 2 Gauss Adaptive 1233 0.768Model 3 MGWR
opt.bandwith.mgwr <- bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWR',
candidates_Kernels = c('bisq','gauss'),
control=list(),
control_search=list()
)mgwr_bisq <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars ='X6',
kernels = c('bisq'),
H = 103,
Model = 'MGWR',
control = list(SE=TRUE,adaptive=TRUE)
)
summary_mgwrsar(mgwr_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("bisq"), H = 103, Model = "MGWR",
#> control = list(SE = TRUE, adaptive = TRUE))
#> Model: MGWR
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 103
#> Computation time: 1.06
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> -0.000223
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 4.78710 -0.09184 -0.02979 -0.01775 -0.03303 -0.07
#> 1st Qu. 10.43257 -0.05408 0.04295 0.00330 0.02346 -0.04
#> Median 12.74609 -0.01505 0.06399 0.00965 0.09408 -0.03
#> Mean 12.46308 -0.02195 0.06648 0.00795 0.09691 -0.03
#> 3rd Qu. 14.25645 0.01239 0.09711 0.01312 0.16275 -0.02
#> Max. 21.27719 0.05583 0.16186 0.02187 0.25029 0.05
#> Effective degrees of freedom: 448
#> AICc: 1446
#> Residual sum of squares: 269
#> RMSE: 0.723mgwr_gauss <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars ='X6',
kernels = c('gauss'),
H = 43 ,
Model = 'MGWR',
control = list(SE=TRUE,adaptive=TRUE)
)
summary_mgwrsar(mgwr_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("gauss"), H = 43, Model = "MGWR",
#> control = list(SE = TRUE, adaptive = TRUE))
#> Model: MGWR
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 43
#> Computation time: 1.21
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> 9.11e-05
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 5.61123 -0.09657 0.01108 -0.00561 -0.03024 -0.05
#> 1st Qu. 9.29834 -0.04103 0.05541 0.00465 0.02823 -0.04
#> Median 11.77222 -0.02127 0.07842 0.01051 0.09551 -0.03
#> Mean 11.52179 -0.02746 0.07874 0.00929 0.09130 -0.03
#> 3rd Qu. 13.75360 -0.00345 0.10194 0.01392 0.14794 -0.02
#> Max. 18.01122 0.02540 0.14723 0.02374 0.18815 0.00
#> Effective degrees of freedom: 468
#> AICc: 1423
#> Residual sum of squares: 300
#> RMSE: 0.763Model_MGWR <- c("Bisquare Adaptive", "Gauss Adaptive")
AIC_MGWR <- c(1191.419, 1226.738)
RMSE_MGWR <- c(mgwr_bisq$RMSE, mgwr_gauss$RMSE)
data.frame(Model_MGWR, AIC_MGWR, RMSE_MGWR)
#> Model_MGWR AIC_MGWR RMSE_MGWR
#> 1 Bisquare Adaptive 1191 0.723
#> 2 Gauss Adaptive 1227 0.763opt.bandwith.mgwr2 <- bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord=coord,
fixed_vars='X6',
Models='MGWR',
candidates_Kernels=c('bisq','gauss'),
control=list(),control_search=list()
)mgwr_bisq2 <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord=coord,
fixed_vars ='X6',
kernels=c('bisq'),
H=65 ,
Model = 'MGWR',
control=list(SE=TRUE,adaptive=TRUE)
)
summary_mgwrsar(mgwr_bisq2)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("bisq"), H = 65, Model = "MGWR",
#> control = list(SE = TRUE, adaptive = TRUE))
#> Model: MGWR
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 65
#> Computation time: 1.05
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> -0.00172
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. -0.894978 -0.116806 -0.055643 -0.041867 -0.117087 -0.12
#> 1st Qu. 9.565599 -0.050794 0.042307 0.000196 0.020622 -0.05
#> Median 12.246055 -0.013896 0.077965 0.007129 0.100859 -0.03
#> Mean 12.158480 -0.019608 0.073614 0.007163 0.098491 -0.03
#> 3rd Qu. 13.896620 0.025971 0.105045 0.014957 0.183649 -0.01
#> Max. 25.061299 0.094110 0.215705 0.032334 0.256565 0.07
#> Effective degrees of freedom: 416
#> AICc: 1493
#> Residual sum of squares: 225
#> RMSE: 0.662mgwr_gauss2 <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'X6',
kernels = c('gauss'),
H = 28 ,
Model = 'MGWR',
control = list(SE=TRUE,adaptive=TRUE))
summary_mgwrsar(mgwr_gauss2)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("gauss"), H = 28, Model = "MGWR",
#> control = list(SE = TRUE, adaptive = TRUE))
#> Model: MGWR
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 28
#> Computation time: 1.16
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> -0.00142
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 4.60340 -0.12346 -0.01526 -0.01028 -0.05372 -0.07
#> 1st Qu. 9.08119 -0.05151 0.06166 0.00338 0.02972 -0.04
#> Median 11.93396 -0.01652 0.07597 0.00905 0.09277 -0.02
#> Mean 11.67532 -0.02695 0.07864 0.00981 0.09173 -0.03
#> 3rd Qu. 13.23546 0.00728 0.10293 0.01449 0.16123 -0.02
#> Max. 21.00384 0.04309 0.14808 0.04155 0.20635 0.01
#> Effective degrees of freedom: 449
#> AICc: 1443
#> Residual sum of squares: 267
#> RMSE: 0.721Model_MGWR2 <- c("Bisquare Adaptive", "Gauss Adaptive")
AIC_MGWR2 <- c(1191.419, 1226.738)
RMSE_MGWR2 <- c(mgwr_bisq2$RMSE, mgwr_gauss2$RMSE)
data.frame(Model_MGWR2, AIC_MGWR2, RMSE_MGWR2)
#> Model_MGWR2 AIC_MGWR2 RMSE_MGWR2
#> 1 Bisquare Adaptive 1191 0.662
#> 2 Gauss Adaptive 1227 0.721Model 4 SAR (MGWRSAR (0,k,0))
opt.bandwith.mgwrsar_0_kc_0 <- bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = '',
candidates_Kernels = c('bisq','gauss'),
control=list(),
control_search=list())mgwrsar_0_kc_0_bisq <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars ='Intercept', kernels=c('bisq'),H=103, Model = 'MGWRSAR_0_kc_0',
control=list(SE=TRUE,adaptive=TRUE))
summary_mgwrsar(mgwrsar_0_kc_0_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "Intercept", kernels = c("bisq"), H = 103, Model = "MGWRSAR_0_kc_0",
#> control = list(SE = TRUE, adaptive = TRUE))
#> Model: MGWRSAR_0_kc_0
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 103
#> Computation time: 1.06
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: Intercept
#> 11.9
#> Varying parameters: X1 X5 X6 X7 X8 X9
#> X1 X5 X6 X7 X8 X9
#> Min. -0.086904 -0.003411 -0.025351 -0.023243 -0.028506 -0.07
#> 1st Qu. -0.047410 0.050616 -0.001617 0.000892 0.029043 -0.04
#> Median -0.007679 0.077588 0.003024 0.008500 0.107882 -0.03
#> Mean -0.016276 0.068839 0.004473 0.006675 0.101186 -0.03
#> 3rd Qu. 0.015483 0.084611 0.007075 0.012707 0.161377 -0.01
#> Max. 0.071022 0.122298 0.035319 0.033022 0.221750 0.05
#> Effective degrees of freedom: 448
#> AICc: 1448
#> Residual sum of squares: 271
#> RMSE: 0.725mgwrsar_0_kc_0_gauss <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
kernels=c('gauss'),
H = 43,
Model = 'MGWRSAR_0_kc_0',
control = list(SE=TRUE,adaptive=TRUE))
summary_mgwrsar(mgwrsar_0_kc_0_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "Intercept", kernels = c("gauss"), H = 43, Model = "MGWRSAR_0_kc_0",
#> control = list(SE = TRUE, adaptive = TRUE))
#> Model: MGWRSAR_0_kc_0
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 43
#> Computation time: 1.19
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: Intercept
#> 11.8
#> Varying parameters: X1 X5 X6 X7 X8 X9
#> X1 X5 X6 X7 X8 X9
#> Min. -0.09488 0.03946 -0.00876 -0.01032 -0.01681 -0.05
#> 1st Qu. -0.04237 0.06365 -0.00105 0.00352 0.02989 -0.04
#> Median -0.02031 0.07570 0.00184 0.00953 0.10888 -0.03
#> Mean -0.02518 0.07227 0.00359 0.00740 0.09543 -0.03
#> 3rd Qu. -0.00144 0.08143 0.00578 0.01355 0.14813 -0.02
#> Max. 0.03543 0.09948 0.02945 0.01990 0.19489 0.00
#> Effective degrees of freedom: 468
#> AICc: 1423
#> Residual sum of squares: 300
#> RMSE: 0.764Model 5 MGWR SAR (0,0,k)
Varying spatial Beta
Bisquare
Optimum Bandwith
opt.bandwith.mgwr00kv_bisq <- bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_0_0_kv',
candidates_Kernels = c('bisq'),
control = list(W=W_bisq),control_search=list())Model
mgwrsar_0_0_kv_bisq <-
MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = NULL,
kernels = c('bisq'),
H = 185, Model = 'MGWRSAR_0_0_kv',
control=list(SE=TRUE,adaptive=TRUE,W=W_bisq))
summary_mgwrsar(mgwrsar_0_0_kv_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = NULL, kernels = c("bisq"), H = 185, Model = "MGWRSAR_0_0_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_bisq))
#> Model: MGWRSAR_0_0_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 185
#> Computation time: 1.11
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: lambda
#> 0.151
#> Varying parameters: Intercept X1 X5 X6 X7 X8 X9
#> Intercept X1 X5 X6 X7 X8
#> Min. 4.567613 -0.072848 0.026195 -0.011132 -0.007224 0.002891
#> 1st Qu. 8.080914 -0.035640 0.043747 0.000607 0.005093 0.023830
#> Median 9.790216 -0.015037 0.056088 0.001997 0.008590 0.112761
#> Mean 9.818628 -0.016831 0.059459 0.003931 0.008011 0.097877
#> 3rd Qu. 11.982679 0.001180 0.075360 0.005267 0.013689 0.132422
#> Max. 12.877866 0.044964 0.102358 0.022017 0.022278 0.216075
#> X9
#> Min. -0.04
#> 1st Qu. -0.03
#> Median -0.03
#> Mean -0.02
#> 3rd Qu. -0.01
#> Max. 0.03
#> Effective degrees of freedom: 469
#> AICc: 1421
#> Residual sum of squares: 300
#> RMSE: 0.764Gauss
Optimum Bandwith
opt.bandwith.mgwr00kv_gauss <- bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_0_0_kv',
candidates_Kernels = c('gauss'),
control = list(W = W_gauss),
control_search = list()
)Model
mgwrsar_0_0_kv_gauss <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = NULL,
kernels = c('bisq'),
H = 48,
Model = 'MGWRSAR_0_0_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_gauss)
)
summary_mgwrsar(mgwrsar_0_0_kv_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = NULL, kernels = c("bisq"), H = 48, Model = "MGWRSAR_0_0_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_gauss))
#> Model: MGWRSAR_0_0_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 48
#> Computation time: 1.01
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: lambda
#> 0.124
#> Varying parameters: Intercept X1 X5 X6 X7 X8 X9
#> Intercept X1 X5 X6 X7 X8
#> Min. -7.261364 -0.140791 -0.144319 -0.043699 -0.063903 -0.198192
#> 1st Qu. 6.725218 -0.058143 0.019069 -0.007248 -0.003641 0.019718
#> Median 10.340695 -0.010109 0.071800 -0.000297 0.005213 0.104157
#> Mean 10.293243 -0.011495 0.068712 -0.000182 0.005737 0.102973
#> 3rd Qu. 14.011053 0.032179 0.110430 0.008624 0.019196 0.181969
#> Max. 28.803334 0.200646 0.326328 0.034703 0.042406 0.312693
#> X9
#> Min. -0.17
#> 1st Qu. -0.05
#> Median -0.03
#> Mean -0.03
#> 3rd Qu. 0.00
#> Max. 0.11
#> Effective degrees of freedom: 367
#> AICc: 1624
#> Residual sum of squares: 194
#> RMSE: 0.614Model_MGWRSAR00k <- c("Bisquare Adaptive", "Gauss Adaptive")
AIC_MGWRSAR00k <- c(1226.462, 1104.081)
RMSE_MGWRSAR00k <- c(mgwrsar_0_0_kv_bisq$RMSE, mgwrsar_0_0_kv_gauss$RMSE)
data.frame(Model_MGWRSAR00k, AIC_MGWRSAR00k, RMSE_MGWRSAR00k)
#> Model_MGWRSAR00k AIC_MGWRSAR00k RMSE_MGWRSAR00k
#> 1 Bisquare Adaptive 1226 0.764
#> 2 Gauss Adaptive 1104 0.614Model 6 MGWR SAR (1,0,kv)
Bisquare
Optimum Bandwith
opt.bandwith.mgwr10kv_bisq <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_1_0_kv',
candidates_Kernels = c('bisq'),
control = list(W = W_bisq),
control_search = list()
)mgwrsar_1_0_kv_bisq <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = NULL,
kernels = c('bisq'),
H = 270,
Model = 'MGWRSAR_1_0_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_bisq)
)
summary_mgwrsar(mgwrsar_1_0_kv_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = NULL, kernels = c("bisq"), H = 270, Model = "MGWRSAR_1_0_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_bisq))
#> Model: MGWRSAR_1_0_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 270
#> Computation time: 0.65
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Varying parameters: Intercept X1 X5 X6 X7 X8 X9
#> Intercept X1 X5 X6 X7 X8
#> Min. 2.99e+00 -5.70e-02 2.20e-02 -6.19e-03 -1.74e-03 1.54e-02
#> 1st Qu. 5.14e+00 -2.68e-02 2.96e-02 7.07e-04 3.53e-03 5.33e-02
#> Median 7.03e+00 -7.89e-03 5.51e-02 1.96e-03 8.36e-03 1.19e-01
#> Mean 6.73e+00 -1.27e-02 5.52e-02 2.93e-03 7.91e-03 9.99e-02
#> 3rd Qu. 7.88e+00 -5.43e-05 6.50e-02 3.62e-03 1.21e-02 1.32e-01
#> Max. 1.21e+01 3.00e-02 1.19e-01 1.52e-02 1.54e-02 1.78e-01
#> X9 lambda
#> Min. -3.01e-02 -0.10
#> 1st Qu. -2.47e-02 0.36
#> Median -2.21e-02 0.39
#> Mean -1.74e-02 0.34
#> 3rd Qu. -1.20e-02 0.41
#> Max. 6.68e-03 0.53
#> Effective degrees of freedom: 479
#> AICc: 1402
#> Residual sum of squares: 309
#> RMSE: 0.775Gauss
Optimum Bandwith
opt.bandwith.mgwr10kv_gauss <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_1_0_kv',
candidates_Kernels = c('gauss'),
control = list(W = W_gauss),
control_search = list()
)mgwrsar_1_0_kv_gauss <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = NULL,
kernels = c('gauss'),
H = 59,
Model = 'MGWRSAR_1_0_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_gauss)
)
summary_mgwrsar(mgwrsar_1_0_kv_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = NULL, kernels = c("gauss"), H = 59, Model = "MGWRSAR_1_0_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_gauss))
#> Model: MGWRSAR_1_0_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 59
#> Computation time: 0.72
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Varying parameters: Intercept X1 X5 X6 X7 X8 X9
#> Intercept X1 X5 X6 X7 X8
#> Min. 2.404756 -0.072267 -0.023757 -0.002830 -0.005969 -0.019615
#> 1st Qu. 4.958718 -0.031364 0.032609 0.000512 0.004867 0.027758
#> Median 7.446119 -0.015179 0.055526 0.002747 0.008830 0.093242
#> Mean 7.257700 -0.019958 0.055479 0.003669 0.007716 0.083996
#> 3rd Qu. 9.400885 -0.003406 0.071246 0.005109 0.011544 0.129913
#> Max. 13.599547 0.019014 0.132030 0.022778 0.015054 0.176316
#> X9 lambda
#> Min. -0.041960 -0.13
#> 1st Qu. -0.029663 0.19
#> Median -0.023150 0.32
#> Mean -0.021123 0.33
#> 3rd Qu. -0.012399 0.44
#> Max. 0.000994 0.88
#> Effective degrees of freedom: 470
#> AICc: 1418
#> Residual sum of squares: 301
#> RMSE: 0.765Model_MGWRSAR_1_0_kv <- c("Bisquare Adaptive", "Gauss Adaptive")
AIC_MGWRSAR_1_0_kv <- c(1230.931 , 1226.788)
RMSE_MGWRSAR_1_0_kv <- c(mgwrsar_1_0_kv_bisq$RMSE, mgwrsar_1_0_kv_gauss$RMSE)
data.frame(Model_MGWRSAR_1_0_kv, AIC_MGWRSAR_1_0_kv, RMSE_MGWRSAR_1_0_kv)
#> Model_MGWRSAR_1_0_kv AIC_MGWRSAR_1_0_kv RMSE_MGWRSAR_1_0_kv
#> 1 Bisquare Adaptive 1231 0.775
#> 2 Gauss Adaptive 1227 0.765Model 7 MGWR SAR (0,kc,kv)
Bisquare
Optimum Bandwith
opt.bandwith.mgwr0kckv_bisq <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_0_kc_kv',
candidates_Kernels = c('bisq'),
control = list(W = W_bisq),
control_search = list()
)Model
mgwrsar_0_kc_kv_bisq <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'X6',
kernels = c('bisq'),
H = 106,
Model = 'MGWRSAR_0_kc_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_bisq)
)
summary_mgwrsar(mgwrsar_0_kc_kv_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("bisq"), H = 106, Model = "MGWRSAR_0_kc_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_bisq))
#> Model: MGWRSAR_0_kc_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 106
#> Computation time: 1.08
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6 lambda
#> -0.000711 0.134
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 3.46999 -0.08656 -0.03182 -0.01769 -0.02966 -0.07
#> 1st Qu. 8.62885 -0.05156 0.03311 0.00344 0.02226 -0.04
#> Median 11.03878 -0.01302 0.05782 0.00974 0.08824 -0.03
#> Mean 10.69843 -0.01976 0.05997 0.00798 0.09409 -0.03
#> 3rd Qu. 12.85812 0.01308 0.09002 0.01306 0.15669 -0.02
#> Max. 18.72866 0.05550 0.13639 0.02262 0.24539 0.05
#> Effective degrees of freedom: 448
#> AICc: 1441
#> Residual sum of squares: 266
#> RMSE: 0.719Gauss
Optimum Bandwith
opt.bandwith.mgwr0kckv_gauss <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_0_kc_kv',
candidates_Kernels = c('gauss'),
control = list(W = W_gauss),
control_search = list()
)Model
mgwrsar_0_kc_kv_gauss <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'X6',
kernels = c('gauss'),
H = 35,
Model = 'MGWRSAR_0_kc_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_gauss)
)
summary_mgwrsar(mgwrsar_0_kc_kv_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("gauss"), H = 35, Model = "MGWRSAR_0_kc_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_gauss))
#> Model: MGWRSAR_0_kc_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 35
#> Computation time: 1.18
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6 lambda
#> -0.00102 0.123
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 4.58574 -0.10806 -0.00257 -0.00917 -0.04032 -0.06
#> 1st Qu. 7.83228 -0.04249 0.04947 0.00449 0.02959 -0.04
#> Median 10.32955 -0.01665 0.07016 0.00972 0.08395 -0.03
#> Mean 10.13883 -0.02533 0.07092 0.00938 0.08720 -0.03
#> 3rd Qu. 12.08595 0.00345 0.09377 0.01355 0.14673 -0.02
#> Max. 17.46650 0.02836 0.13273 0.03197 0.19396 0.00
#> Effective degrees of freedom: 458
#> AICc: 1427
#> Residual sum of squares: 277
#> RMSE: 0.734Model_MGWRSAR_0_kc_kv <- c("Bisquare Adaptive", "Gauss Adaptive")
AIC_MGWRSAR_0_kc_kv <- c(1184.993, 1195.771)
RMSE_MGWRSAR_0_kc_kv <- c(mgwrsar_0_kc_kv_bisq$RMSE, mgwrsar_0_kc_kv_gauss$RMSE)
data.frame(Model_MGWRSAR_0_kc_kv, AIC_MGWRSAR_0_kc_kv, RMSE_MGWRSAR_0_kc_kv)
#> Model_MGWRSAR_0_kc_kv AIC_MGWRSAR_0_kc_kv RMSE_MGWRSAR_0_kc_kv
#> 1 Bisquare Adaptive 1185 0.719
#> 2 Gauss Adaptive 1196 0.734Model 8 MGWR SAR (1,kc,kv)
Bisquare
Optimum Bandwidth
opt.bandwith.mgwr1kckv_bisq <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_1_kc_kv',
candidates_Kernels = c('bisq'),
control = list(W = W_bisq),
control_search = list()
)Model
# Model
mgwrsar_1_kc_kv_bisq <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'X6',
kernels = c('bisq'),
H = 177,
Model = 'MGWRSAR_1_kc_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_bisq)
)
summary_mgwrsar(mgwrsar_1_kc_kv_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("bisq"), H = 177, Model = "MGWRSAR_1_kc_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_bisq))
#> Model: MGWRSAR_1_kc_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 177
#> Computation time: 1.44
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> 0.000691
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 1.27950 -0.07166 -0.00918 -0.00409 0.00169 -0.04592
#> 1st Qu. 7.46963 -0.03568 0.02880 0.00398 0.02372 -0.03440
#> Median 9.10806 -0.02178 0.04660 0.00949 0.09811 -0.02864
#> Mean 9.58946 -0.01899 0.05424 0.00844 0.09271 -0.02442
#> 3rd Qu. 12.16567 0.00337 0.08003 0.01347 0.13055 -0.01616
#> Max. 20.68604 0.04078 0.14569 0.01826 0.20821 0.02772
#> lambda
#> Min. -0.45
#> 1st Qu. 0.08
#> Median 0.24
#> Mean 0.21
#> 3rd Qu. 0.38
#> Max. 0.60
#> Effective degrees of freedom: 467
#> AICc: 1424
#> Residual sum of squares: 299
#> RMSE: 0.763Gauss
Optimum Bandwidth
opt.bandwith.mgwr1kckv_gauss <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_1_kc_kv',
candidates_Kernels = c('gauss'),
control = list(W = W_gauss),
control_search = list()
)
#>
#> ..........Model
# Model
mgwrsar_1_kc_kv_gauss <-
MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'X6',
kernels = c('gauss'),
H = 511,
Model = 'MGWRSAR_1_kc_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_gauss)
)
summary_mgwrsar(mgwrsar_1_kc_kv_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("gauss"), H = 511, Model = "MGWRSAR_1_kc_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_gauss))
#> Model: MGWRSAR_1_kc_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 511
#> Computation time: 1.64
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> -0.00133
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 3.76127 -0.02183 0.09342 0.00855 0.02997 -0.02678
#> 1st Qu. 3.85384 -0.02125 0.09367 0.00891 0.03381 -0.02638
#> Median 3.91847 -0.02036 0.09450 0.00916 0.03669 -0.02603
#> Mean 3.91057 -0.02037 0.09627 0.00903 0.03535 -0.02601
#> 3rd Qu. 3.97163 -0.01943 0.09914 0.00918 0.03700 -0.02563
#> Max. 4.01050 -0.01915 0.10210 0.00920 0.03714 -0.02533
#> lambda
#> Min. 0.38
#> 1st Qu. 0.40
#> Median 0.41
#> Mean 0.41
#> 3rd Qu. 0.42
#> Max. 0.42
#> Effective degrees of freedom: 504
#> AICc: 1382
#> Residual sum of squares: 352
#> RMSE: 0.828Model 9 MGWR SAR (1,kc,0)
Bisquare
Optimum Bandwith
opt.bandwith.mgwr1kc0_bisq <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_1_kc_0',
candidates_Kernels = c('bisq'),
control = list(W = W_bisq),
control_search = list()
)Model
Fixed X6
mgwrsar_1_kc_kv_bisq <- MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'X6',
kernels = c('bisq'),
H = 231 ,
Model = 'MGWRSAR_1_kc_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_bisq)
)
summary_mgwrsar(mgwrsar_1_kc_kv_bisq)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("bisq"), H = 231, Model = "MGWRSAR_1_kc_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_bisq))
#> Model: MGWRSAR_1_kc_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: bisq
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 231
#> Computation time: 1.39
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> 0.00166
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 3.922328 -0.064083 0.007483 -0.000366 0.014755 -0.035630
#> 1st Qu. 6.074547 -0.028612 0.036410 0.003451 0.048327 -0.029965
#> Median 8.652182 -0.016881 0.052551 0.008861 0.109999 -0.025452
#> Mean 8.371140 -0.015839 0.056568 0.008240 0.100042 -0.021200
#> 3rd Qu. 10.175450 0.001590 0.068968 0.012767 0.131166 -0.015083
#> Max. 17.509189 0.034055 0.134258 0.014028 0.196421 0.017631
#> lambda
#> Min. -0.24
#> 1st Qu. 0.21
#> Median 0.27
#> Mean 0.26
#> 3rd Qu. 0.40
#> Max. 0.59
#> Effective degrees of freedom: 477
#> AICc: 1408
#> Residual sum of squares: 310
#> RMSE: 0.777Gauss
Optimum Bandwith
opt.bandwith.mgwr1kc0_gauss <-
bandwidths_mgwrsar(
formula = 'log(Y)~X1+X5+X6+X7+X8+X9',
data = datgwr2,
coord = coord,
fixed_vars = 'Intercept',
Models = 'MGWRSAR_1_kc_0',
candidates_Kernels = c('gauss'),
control = list(W = W_gauss),
control_search = list()
)Model
Fixed X6
mgwrsar_1_kc_kv_gauss <-
MGWRSAR(
formula = 'log(Y)~.',
data = datgwr2,
coord = coord,
fixed_vars = 'X6',
kernels = c('gauss'),
H = 311 ,
Model = 'MGWRSAR_1_kc_kv',
control = list(SE = TRUE, adaptive = TRUE, W = W_gauss)
)
summary_mgwrsar(mgwrsar_1_kc_kv_gauss)
#> Call:
#> MGWRSAR(formula = "log(Y)~.", data = datgwr2, coords = coord,
#> fixed_vars = "X6", kernels = c("gauss"), H = 311, Model = "MGWRSAR_1_kc_kv",
#> control = list(SE = TRUE, adaptive = TRUE, W = W_gauss))
#> Model: MGWRSAR_1_kc_kv
#> Method for spatial autocorrelation: 2SLS
#> Kernels function: gauss
#> Kernels adaptive: YES
#> Kernels type: GD
#> Bandwidth: 311
#> Computation time: 1.67
#> Use of parallel computing: FALSE ncore = 1
#> Use of rough kernel: NO
#> Use of Target Points: NO
#> Number of data points: 514
#> Constant parameters: X6
#> -0.000347
#> Varying parameters: Intercept X1 X5 X7 X8 X9
#> Intercept X1 X5 X7 X8 X9
#> Min. 3.02971 -0.02506 0.06714 0.00793 0.02793 -0.02715
#> 1st Qu. 3.40246 -0.01942 0.07162 0.00870 0.04014 -0.02541
#> Median 4.04258 -0.01681 0.07689 0.00912 0.07111 -0.02058
#> Mean 3.97869 -0.01800 0.08407 0.00891 0.06554 -0.02126
#> 3rd Qu. 4.44589 -0.01522 0.10031 0.00924 0.08765 -0.01829
#> Max. 4.99890 -0.01381 0.10711 0.00945 0.10067 -0.01483
#> lambda
#> Min. 0.37
#> 1st Qu. 0.41
#> Median 0.43
#> Mean 0.43
#> 3rd Qu. 0.44
#> Max. 0.48
#> Effective degrees of freedom: 501
#> AICc: 1379
#> Residual sum of squares: 339
#> RMSE: 0.813Model_GWRSAR_1_kc_kv <- c("Bisquare Adaptive", "Gauss Adaptive")
AIC_GWRSAR_1_kc_kv <- c(1234.797, 1257.486)
RMSE_GWRSAR_1_kc_kv <- c(mgwrsar_1_kc_kv_bisq$RMSE, mgwrsar_1_kc_kv_gauss$RMSE)
data.frame(Model_GWRSAR_1_kc_kv, AIC_GWRSAR_1_kc_kv, RMSE_GWRSAR_1_kc_kv)
#> Model_GWRSAR_1_kc_kv AIC_GWRSAR_1_kc_kv RMSE_GWRSAR_1_kc_kv
#> 1 Bisquare Adaptive 1235 0.777
#> 2 Gauss Adaptive 1257 0.813Interpretasi Model Terbaik
Namun demikian, meski X6 memiliki proporsi dugaan parameter di dalam SK 95% tertinggi, nilai tersebut tidak mencapai proporsi acuan yaitu sebesar 60% untuk dapat dikatakan dugaan parameter bersifat global. Berdasarkan hal tersebut serta pemodelan dengan metode SAR dapat disimpulkan bahwa dugaan parameter dari pemodelan tingkat PDRB kabupaten/kota di Indonesia bersifat lokal disertai adanya dependensi spasial.
Dengan demikian pemodelan yang dipilih untuk dibandingkan sebagai model terbaik adalah dengan menggunakan model MGWR-SAR dengan komponen peubah bersifat lokal yaitu model 5 MGWR-SAR(0,0,6) dan Model 6 MGWR - SAR (1,0,6).
Model yang dibandingkan hanya Berdasarkan kriteria RMSE dalam melihat performa model, terlihat bahwa model MGWR-SAR (0,0,6) dengan pembobot kernel gauss adaptive memiliki nilai RMSE terkecil, sehingga dipilih sebagai model terbaik.
Betav
matrix of coefficients of dim(n,kv) x kv.
Betac
vector of coefficients of length kc = lambda = 0.1243248
beta <- mgwrsar_0_0_kv_gauss$Betav
beta
#> Intercept X1 X5 X6 X7 X8
#> [1,] 2.1959 -0.059379 0.182577 -0.003091 0.015758 0.022109
#> [2,] 7.8320 -0.104108 0.130546 -0.005909 0.015529 -0.021779
#> [3,] 3.7197 -0.077428 0.173292 -0.004315 0.019954 -0.012096
#> [4,] 4.6728 -0.087449 0.166164 -0.004583 0.020602 -0.022967
#> [5,] 4.5238 -0.061401 0.128415 -0.013503 0.015550 0.122117
#> [6,] 4.7810 -0.070556 0.134358 -0.009929 0.019965 0.068257
#> [7,] 3.8525 -0.060494 0.139582 -0.011470 0.017473 0.102139
#> [8,] 4.3379 -0.058009 0.128725 -0.013920 0.014174 0.135539
#> [9,] 4.3330 -0.060504 0.130848 -0.013174 0.015744 0.120784
#> [10,] 5.5064 -0.071217 0.120443 -0.012413 0.018731 0.091698
#> [11,] 5.8000 -0.075361 0.119930 -0.011217 0.020794 0.071013
#> [12,] 3.9909 -0.079739 0.173018 -0.003855 0.017998 -0.015071
#> [13,] 4.7461 -0.077222 0.148945 -0.006155 0.022519 0.011123
#> [14,] 6.9150 -0.091140 0.124684 -0.005711 0.025656 -0.010418
#> [15,] 3.6269 -0.062893 0.147550 -0.009774 0.019224 0.076071
#> [16,] 3.9160 -0.057262 0.135005 -0.012627 0.015567 0.124322
#> [17,] 5.3730 -0.073169 0.125552 -0.010800 0.020191 0.072535
#> [18,] 4.6543 -0.063379 0.127890 -0.013126 0.016333 0.114358
#> [19,] 4.3174 -0.057204 0.128454 -0.014058 0.013769 0.139293
#> [20,] 4.5335 -0.058093 0.125800 -0.014530 0.013313 0.141661
#> [21,] 7.1181 -0.089758 0.118094 -0.006412 0.025467 0.001393
#> [22,] 5.8878 -0.074392 0.116933 -0.012014 0.019837 0.081932
#> [23,] 8.0868 -0.105755 0.127521 -0.005374 0.017056 -0.023484
#> [24,] 13.2191 0.035496 0.059719 -0.001233 0.014835 0.174212
#> [25,] 26.7196 -0.005881 -0.121977 0.014781 0.002959 -0.045537
#> [26,] 19.3635 -0.101333 -0.000531 -0.001984 0.020598 -0.104112
#> [27,] 13.0372 -0.110455 0.071808 -0.005003 0.019114 -0.059076
#> [28,] 14.7414 -0.116966 0.053743 -0.004279 0.021278 -0.067747
#> [29,] 14.8308 -0.123055 0.052054 -0.003817 0.021454 -0.061295
#> [30,] 17.6766 -0.092100 0.025240 -0.001115 0.020092 -0.087017
#> [31,] 14.9169 -0.116302 0.049730 -0.001998 0.019624 -0.055600
#> [32,] 14.3909 -0.135859 0.052358 -0.003068 0.020710 -0.040162
#> [33,] 9.0111 -0.112319 0.115606 -0.005024 0.018666 -0.024471
#> [34,] 8.8320 -0.109633 0.116791 -0.004591 0.019693 -0.026358
#> [35,] 11.2319 -0.127751 0.083728 -0.004288 0.022546 -0.024630
#> [36,] 8.2390 -0.103877 0.116118 -0.004827 0.024889 -0.025774
#> [37,] 14.9250 -0.091440 0.027063 -0.009009 0.005235 -0.060356
#> [38,] 11.5643 -0.115173 0.089052 -0.005481 0.018516 -0.044587
#> [39,] 9.6750 -0.113821 0.109070 -0.005660 0.017774 -0.028102
#> [40,] 12.4305 -0.125856 0.078305 -0.005149 0.020136 -0.041818
#> [41,] 13.2988 -0.139877 0.060165 -0.003740 0.021961 -0.026964
#> [42,] 14.2727 -0.123867 0.054006 -0.002046 0.019356 -0.042144
#> [43,] 21.4772 -0.087723 -0.028464 0.001760 0.017522 -0.107153
#> [44,] 27.4268 0.035448 -0.102955 0.016477 -0.000506 -0.108041
#> [45,] 21.0565 -0.089060 -0.015850 0.002602 0.016696 -0.103541
#> [46,] 17.4947 -0.108800 0.023895 -0.001967 0.021557 -0.079939
#> [47,] 8.4408 -0.097789 0.116051 -0.007494 0.010091 -0.018793
#> [48,] 10.5368 -0.097272 0.087393 -0.007934 0.009135 -0.032107
#> [49,] 14.0955 -0.108986 0.059951 -0.004864 0.019597 -0.068915
#> [50,] 14.9893 -0.110253 0.049598 -0.001679 0.019246 -0.057076
#> [51,] 13.7659 -0.140791 0.058943 -0.004036 0.021327 -0.034909
#> [52,] 13.6960 -0.138037 0.055712 -0.003171 0.021382 -0.028232
#> [53,] 10.7385 -0.122088 0.089231 -0.004353 0.023250 -0.027194
#> [54,] 9.6122 -0.113517 0.103270 -0.004238 0.022846 -0.027955
#> [55,] 20.8405 -0.082930 -0.026005 0.000106 0.017714 -0.108115
#> [56,] 9.5985 -0.099448 0.102370 -0.007390 0.010962 -0.027976
#> [57,] 20.6242 0.008845 -0.030761 0.004885 -0.014951 0.124990
#> [58,] 13.9730 0.050734 0.112554 -0.005898 -0.032633 0.096398
#> [59,] 18.5756 0.102253 0.074996 0.002835 -0.031596 0.065125
#> [60,] 20.9308 0.103499 0.041360 0.007957 -0.029216 0.054897
#> [61,] 22.4395 0.125826 0.004149 0.012505 -0.023395 0.066835
#> [62,] 21.8538 0.164911 0.010503 0.011553 -0.021559 0.085345
#> [63,] 24.3273 0.200646 -0.034125 0.018703 -0.017055 0.088942
#> [64,] 14.1241 -0.127050 0.055187 -0.002295 0.019599 -0.039936
#> [65,] 27.5974 0.127627 -0.127084 0.025052 -0.003359 0.073756
#> [66,] 15.7148 0.061945 0.088056 -0.002112 -0.027323 0.056788
#> [67,] 18.2772 0.065455 0.043157 0.002856 -0.019378 0.013684
#> [68,] 28.0127 0.095946 -0.144319 0.024692 -0.000614 0.066154
#> [69,] 18.9172 0.127525 0.058094 0.003915 -0.026680 0.082652
#> [70,] 19.9843 0.112595 0.045260 0.006174 -0.026788 0.067883
#> [71,] 20.9680 0.108450 0.032602 0.008370 -0.026621 0.061442
#> [72,] 22.3288 0.156783 0.008027 0.012698 -0.023547 0.079068
#> [73,] 23.6735 0.150495 -0.022939 0.016751 -0.019820 0.073879
#> [74,] 24.4250 0.135778 -0.023375 0.017832 -0.023907 0.067765
#> [75,] 21.4699 0.166487 0.015947 0.010273 -0.021481 0.090201
#> [76,] 23.6116 0.060770 -0.015187 0.012151 -0.018055 0.003433
#> [77,] 20.5372 0.027223 -0.029689 0.009928 0.002239 -0.064894
#> [78,] 5.3066 -0.012577 0.065711 0.006978 0.027159 -0.037989
#> [79,] 24.2233 -0.006601 -0.098211 0.018157 0.011745 -0.051418
#> [80,] 6.7259 -0.012357 0.113294 0.013410 0.001932 0.154049
#> [81,] 27.3021 0.066365 -0.070102 0.021786 -0.018781 0.017313
#> [82,] 26.9360 0.037172 -0.099554 0.023282 -0.004900 -0.043852
#> [83,] 15.7276 -0.049437 -0.023882 0.003577 0.042406 -0.054337
#> [84,] 24.2740 -0.064993 -0.054241 0.008434 0.014920 -0.120651
#> [85,] 6.9968 -0.043833 0.049788 0.001363 0.041782 -0.043589
#> [86,] 28.8033 0.018320 -0.097505 0.021061 -0.007000 -0.031150
#> [87,] 21.4282 -0.053446 -0.027137 0.003903 0.027189 -0.117804
#> [88,] 14.4908 0.045490 0.080272 -0.003756 -0.019348 0.059087
#> [89,] 10.0397 0.029324 0.089255 -0.001277 0.000927 0.021938
#> [90,] 9.2718 0.041151 0.129363 -0.003647 0.020544 -0.078050
#> [91,] 8.6831 0.041274 0.144875 -0.003344 0.028025 -0.159370
#> [92,] -0.8460 0.048739 0.280362 -0.026582 0.040550 -0.166430
#> [93,] -2.7079 0.010609 0.247190 -0.026424 0.037063 -0.082822
#> [94,] -2.8393 -0.000817 0.233530 -0.023572 0.036732 -0.078410
#> [95,] 17.3296 0.034136 0.011073 0.004762 0.004359 -0.077883
#> [96,] 14.8389 0.038343 0.047659 -0.000301 -0.006986 0.000871
#> [97,] 1.7165 0.055139 0.259545 -0.020109 0.039427 -0.198192
#> [98,] 11.8555 0.034600 0.093945 -0.006016 -0.017895 0.083484
#> [99,] -0.3692 0.021078 0.263115 -0.028449 0.007931 0.186078
#> [100,] -2.1941 0.085864 0.279162 -0.037237 0.010978 0.184306
#> [101,] 0.3877 0.029575 0.273542 -0.028275 0.012817 0.105770
#> [102,] 0.7081 0.024797 0.263833 -0.027181 0.012014 0.120171
#> [103,] 6.5763 0.060173 0.203369 -0.017345 0.022455 -0.046884
#> [104,] 7.7849 0.079583 0.233344 -0.021692 0.034062 -0.142147
#> [105,] 5.1290 0.102438 0.230874 -0.023792 0.023155 -0.023047
#> [106,] 2.2938 0.006307 0.201710 -0.021822 0.003793 0.245171
#> [107,] -2.2990 0.053753 0.282461 -0.033808 0.008282 0.207996
#> [108,] -2.2850 0.067811 0.326328 -0.037653 0.011610 0.150393
#> [109,] 2.4052 0.039319 0.231759 -0.023574 0.014083 0.088535
#> [110,] 0.9876 0.057237 0.305537 -0.030900 0.017687 0.028519
#> [111,] 7.1694 0.057113 0.186113 -0.013683 0.021916 -0.052959
#> [112,] 1.7329 0.093890 0.271601 -0.030306 0.017040 0.051315
#> [113,] -0.3259 0.043900 0.324207 -0.033163 0.011217 0.108204
#> [114,] 1.0518 0.018273 0.243597 -0.024964 0.010551 0.147229
#> [115,] 4.4242 0.052213 0.214184 -0.020691 0.017626 0.028566
#> [116,] 1.0739 0.018268 0.225905 -0.023185 0.009257 0.169209
#> [117,] 4.1505 0.048391 0.202328 -0.019287 0.015236 0.059150
#> [118,] 4.6071 0.052417 0.186374 -0.017045 0.012147 0.069087
#> [119,] 1.1438 0.008788 0.218404 -0.023218 0.006129 0.220540
#> [120,] 2.4789 0.034186 0.207500 -0.021110 0.011586 0.127566
#> [121,] 9.7731 0.038135 0.109275 -0.009949 -0.011344 0.110497
#> [122,] 2.1263 0.083210 0.287870 -0.029792 0.020268 0.004357
#> [123,] 3.4887 0.047347 0.206788 -0.020892 0.013702 0.084831
#> [124,] 3.7633 0.047131 0.194398 -0.019176 0.012697 0.093255
#> [125,] 3.7237 0.046559 0.189515 -0.018820 0.011826 0.104270
#> [126,] 4.8188 -0.001861 0.149415 -0.012949 -0.002775 0.272058
#> [127,] 6.6530 -0.006175 0.126512 -0.007066 -0.007486 0.255543
#> [128,] 6.1997 -0.047979 0.144671 0.005437 -0.006197 0.187150
#> [129,] 4.9223 -0.043288 0.172560 -0.006018 -0.002527 0.180131
#> [130,] 5.5441 -0.003530 0.146111 -0.012967 -0.001003 0.216359
#> [131,] 4.9922 0.009811 0.150826 -0.012858 -0.002754 0.245605
#> [132,] 2.4552 0.017031 0.197577 -0.020981 0.001812 0.247214
#> [133,] 2.8348 0.030094 0.179298 -0.022655 0.005811 0.208448
#> [134,] 6.1983 -0.028788 0.140608 -0.002108 -0.008433 0.228364
#> [135,] 6.3218 -0.018108 0.134722 -0.004772 -0.007504 0.237764
#> [136,] 0.2665 0.056785 0.221365 -0.029032 0.007734 0.218704
#> [137,] 3.1392 0.024891 0.180748 -0.018839 0.001446 0.230495
#> [138,] 4.1908 -0.002748 0.157104 -0.015146 -0.000749 0.278902
#> [139,] 5.9260 -0.036689 0.148748 -0.001384 -0.006928 0.209617
#> [140,] 5.5467 -0.023496 0.150717 -0.007678 -0.004090 0.214699
#> [141,] 0.4422 0.084211 0.219754 -0.028858 0.021563 0.049335
#> [142,] 9.8874 0.100851 0.026507 -0.001199 0.003888 0.174254
#> [143,] -0.4217 0.081700 0.241124 -0.029645 0.023764 0.013876
#> [144,] 3.5140 0.093321 0.188802 -0.028512 0.017245 0.098892
#> [145,] 6.1296 0.093283 0.155035 -0.027051 0.013896 0.132750
#> [146,] 5.7521 0.078214 0.079236 -0.005223 -0.000191 0.210762
#> [147,] 2.2066 0.091247 0.203154 -0.028443 0.019335 0.074603
#> [148,] -1.8227 -0.063441 0.156366 -0.016511 0.042069 -0.027510
#> [149,] -6.5029 -0.074088 0.208217 -0.033746 0.037203 0.013504
#> [150,] 9.3417 -0.057954 0.004265 -0.017030 0.008294 0.172817
#> [151,] -7.2614 -0.028855 0.209454 -0.029081 0.029398 0.045453
#> [152,] -1.4419 -0.086380 0.169993 -0.043699 0.037503 0.047258
#> [153,] -5.7674 -0.074345 0.203932 -0.029242 0.038720 -0.007054
#> [154,] -6.6112 -0.068361 0.206515 -0.031551 0.036155 0.013094
#> [155,] 6.9585 -0.096750 0.120465 0.004175 0.028499 0.072674
#> [156,] 7.7592 -0.106772 0.084044 0.006931 0.034630 0.083588
#> [157,] 7.7742 -0.113208 0.085606 0.006356 0.035560 0.083914
#> [158,] 7.7193 -0.107105 0.087903 0.006782 0.034737 0.081042
#> [159,] 7.3751 -0.105939 0.086825 0.007071 0.034925 0.086593
#> [160,] 7.8386 -0.109029 0.089858 0.006353 0.035022 0.076980
#> [161,] 6.7451 -0.089206 0.087309 0.008610 0.026257 0.112578
#> [162,] 6.7587 -0.096822 0.051676 0.008535 0.036373 0.113599
#> [163,] 8.1278 -0.112710 0.061992 0.005447 0.032997 0.105380
#> [164,] 5.5084 -0.092486 0.093969 0.001330 0.025637 0.108204
#> [165,] 3.4702 -0.051490 0.101082 -0.010246 0.007703 0.177440
#> [166,] 3.9233 -0.002246 0.066769 -0.019535 -0.002644 0.237983
#> [167,] 9.4423 -0.002807 0.010296 -0.020073 0.003708 0.216810
#> [168,] 10.3585 0.017302 -0.000478 -0.020765 -0.001140 0.220520
#> [169,] 10.8089 0.012857 -0.012734 -0.021914 0.002018 0.225560
#> [170,] 7.4319 -0.020825 0.039056 -0.017450 0.005174 0.208296
#> [171,] 5.5839 -0.072208 0.098884 -0.005031 0.016689 0.128574
#> [172,] 11.3717 -0.054172 0.030511 -0.011226 0.017701 0.137099
#> [173,] 5.5644 -0.087681 0.121488 0.001255 0.027471 0.070715
#> [174,] 6.9112 -0.099733 0.097729 0.005469 0.034269 0.069131
#> [175,] 7.7443 -0.116805 0.087847 0.005088 0.035831 0.079900
#> [176,] 7.9361 -0.118533 0.080300 0.005387 0.035538 0.087478
#> [177,] 5.9826 -0.093698 0.098136 0.003222 0.029016 0.088124
#> [178,] 10.0625 0.017656 0.039347 -0.015709 -0.010003 0.195169
#> [179,] 7.7743 -0.110044 0.067776 0.007056 0.034818 0.099199
#> [180,] 8.1498 -0.115153 0.056651 0.005627 0.032512 0.113863
#> [181,] 5.6585 -0.091983 0.097458 0.001417 0.025689 0.102981
#> [182,] 11.8385 0.018110 -0.022269 -0.021587 0.001389 0.220016
#> [183,] 7.9629 -0.114510 0.084028 0.005925 0.035349 0.082160
#> [184,] 7.6695 -0.110159 0.076194 0.007103 0.035144 0.093470
#> [185,] 5.5563 -0.091994 0.102116 0.002401 0.027628 0.091533
#> [186,] 5.6267 0.008346 0.048634 -0.020426 -0.003501 0.236808
#> [187,] 10.7325 0.019230 0.019056 -0.017470 -0.006555 0.200439
#> [188,] 12.1702 0.024942 0.063839 -0.003717 -0.022161 0.142237
#> [189,] 12.1166 0.017306 0.074757 0.001949 -0.019726 0.106624
#> [190,] 12.5227 0.016279 0.070787 0.000898 -0.020578 0.104045
#> [191,] 14.2503 -0.009933 0.045803 0.005501 -0.009174 0.048203
#> [192,] 12.5476 0.008015 0.065455 0.004262 -0.012752 0.081556
#> [193,] 19.0999 -0.032491 -0.013266 0.007229 -0.006008 0.017725
#> [194,] 17.4585 -0.028536 0.011058 0.006762 -0.009447 0.023125
#> [195,] 15.8973 -0.033677 0.012821 0.006352 -0.004210 0.016136
#> [196,] 9.3485 -0.028313 0.062735 0.004741 0.002957 0.019618
#> [197,] 9.1164 -0.026827 0.062193 0.004615 0.004566 0.019258
#> [198,] 7.3075 -0.024174 0.071423 0.004693 0.008914 0.018377
#> [199,] 7.9543 -0.012074 0.072796 0.002831 0.012392 0.003551
#> [200,] 7.3217 -0.022822 0.069203 0.005221 0.009835 0.018713
#> [201,] 7.2070 -0.026752 0.067163 0.005693 0.008709 0.024381
#> [202,] 7.1135 -0.032325 0.062516 0.006634 0.006510 0.032639
#> [203,] 5.3696 0.003116 0.041477 0.005133 0.007617 0.124051
#> [204,] 5.4636 -0.010286 0.032407 0.007121 0.006475 0.118631
#> [205,] 6.9433 -0.031766 0.048019 0.008498 0.006322 0.048610
#> [206,] 7.4256 -0.035090 0.064637 0.006731 0.004267 0.031455
#> [207,] 8.0836 -0.035293 0.072281 0.006489 0.001372 0.026514
#> [208,] 7.9425 -0.034803 0.073755 0.005709 0.001608 0.025351
#> [209,] 13.1853 -0.031492 0.042143 0.005773 -0.004503 0.020352
#> [210,] 16.3522 -0.036110 0.016402 0.006752 -0.007682 0.015771
#> [211,] 15.8114 -0.039450 0.023278 0.006928 -0.008369 0.012996
#> [212,] 15.3750 -0.034971 0.034640 0.006678 -0.010094 0.016994
#> [213,] 12.4601 -0.004719 0.073959 0.004074 -0.015249 0.060366
#> [214,] 12.8378 0.013021 0.059165 0.000677 -0.018477 0.096854
#> [215,] 13.4510 0.025654 0.023944 -0.003336 -0.014958 0.126424
#> [216,] 12.9449 0.023337 0.012360 -0.011193 -0.009953 0.162419
#> [217,] 16.5237 -0.036510 0.011227 0.006693 -0.006512 0.014586
#> [218,] 7.2869 -0.026862 0.071687 0.004827 0.007809 0.020575
#> [219,] 10.2102 -0.032180 0.059598 0.005107 -0.000378 0.019458
#> [220,] 10.8398 -0.034869 0.061853 0.005614 -0.003252 0.018966
#> [221,] 12.4262 -0.004782 0.073879 0.004131 -0.015010 0.060015
#> [222,] 13.6828 0.027602 0.019099 -0.003776 -0.014584 0.127914
#> [223,] 19.3011 -0.035011 -0.014307 0.007370 -0.006949 0.015006
#> [224,] 15.1932 -0.029810 0.012936 0.005973 -0.000706 0.016922
#> [225,] 9.6682 -0.022575 0.053873 0.004397 0.006957 0.016842
#> [226,] 14.9752 -0.032828 0.018255 0.006106 -0.002684 0.015994
#> [227,] 14.2259 -0.029805 0.021963 0.005771 -0.000621 0.017221
#> [228,] 8.7740 -0.006645 0.074583 -0.002849 0.016038 -0.006179
#> [229,] 8.6025 0.030597 0.087833 -0.014572 0.019931 0.046631
#> [230,] 8.0737 0.074152 0.107141 -0.038593 0.023452 0.214758
#> [231,] 8.4781 0.089678 0.109564 -0.036672 0.015708 0.268160
#> [232,] 10.3984 0.076536 0.103348 -0.005491 0.003126 0.213310
#> [233,] 9.9133 0.075029 0.096723 -0.001948 -0.007642 0.240833
#> [234,] 11.6322 0.065621 0.093470 0.001430 0.002837 0.204125
#> [235,] 13.0362 0.040719 0.066671 -0.000562 0.011933 0.180387
#> [236,] 13.2244 0.035482 0.059763 -0.001208 0.014790 0.174202
#> [237,] 13.7851 0.015324 0.046509 -0.007939 0.021293 0.177961
#> [238,] 12.7597 0.033360 0.062676 -0.001422 0.015441 0.176618
#> [239,] 12.1854 0.036146 0.067238 0.000426 0.013922 0.179083
#> [240,] 11.9421 0.040892 0.068298 0.002930 0.010742 0.183871
#> [241,] 11.6717 0.056659 0.075301 0.005025 0.003262 0.205614
#> [242,] 10.6087 0.063262 0.079024 0.009462 -0.002585 0.224887
#> [243,] 10.1355 0.071758 0.090903 0.009987 -0.008482 0.237421
#> [244,] 8.5403 0.076069 0.095230 0.008917 -0.017785 0.269913
#> [245,] 8.4300 0.080612 0.081550 -0.005428 -0.003897 0.266754
#> [246,] 7.7113 0.070710 0.081425 -0.017972 0.012613 0.204117
#> [247,] 7.7590 0.022647 0.077534 -0.000892 0.014882 0.026275
#> [248,] 6.7938 0.030153 0.063626 0.005668 0.012801 0.053934
#> [249,] 5.8732 0.068823 0.062471 0.010327 -0.006744 0.281812
#> [250,] 4.5718 0.070470 0.073411 0.019736 -0.014269 0.312693
#> [251,] 7.7376 0.074104 0.089280 0.017369 -0.019108 0.280734
#> [252,] 9.0512 0.068373 0.082962 0.014634 -0.009081 0.249280
#> [253,] 9.2845 0.057269 0.074640 0.012567 -0.000833 0.227088
#> [254,] 9.7924 0.049730 0.072083 0.009943 0.004203 0.209393
#> [255,] 10.2947 0.044943 0.072356 0.007589 0.007535 0.196320
#> [256,] 10.2749 0.043901 0.077136 0.006362 0.009444 0.188953
#> [257,] 9.4112 0.079313 0.093403 -0.011480 -0.003345 0.256733
#> [258,] 9.9038 0.079339 0.106800 -0.010531 0.002517 0.226694
#> [259,] 11.5809 0.066232 0.094444 0.001405 0.002657 0.204636
#> [260,] 11.8727 0.045396 0.072246 0.003399 0.008480 0.189905
#> [261,] 11.4842 0.055135 0.072993 0.005963 0.003343 0.206814
#> [262,] 9.4735 0.074479 0.093935 0.011762 -0.012833 0.250532
#> [263,] 8.0669 0.046272 0.075139 -0.007250 0.014930 0.089281
#> [264,] 9.8330 0.064109 0.078553 0.012097 -0.004570 0.234587
#> [265,] 10.8729 0.071593 0.099208 0.003724 -0.001820 0.216535
#> [266,] 6.7471 -0.085033 0.090272 0.009287 0.023094 0.118538
#> [267,] 6.7943 -0.092942 0.071646 0.008629 0.031093 0.111535
#> [268,] 7.2743 -0.098494 0.083491 0.007738 0.032433 0.093592
#> [269,] 6.8623 -0.074992 0.103931 0.010769 0.013687 0.132973
#> [270,] 7.3803 -0.103131 0.088559 0.007084 0.034095 0.085488
#> [271,] 6.9444 -0.072173 0.113460 0.010725 0.009868 0.135564
#> [272,] 6.8372 -0.083685 0.100571 0.008999 0.020576 0.115104
#> [273,] 7.4945 -0.104977 0.080950 0.007433 0.034672 0.089826
#> [274,] 14.6091 0.005190 0.031315 -0.010553 0.023732 0.181478
#> [275,] 15.7002 -0.007789 0.005924 -0.013913 0.028191 0.189275
#> [276,] 15.7816 -0.009001 0.003158 -0.014286 0.028767 0.190679
#> [277,] 15.8117 -0.010513 -0.001607 -0.014729 0.029619 0.195635
#> [278,] 15.7807 -0.010505 -0.002660 -0.014735 0.029488 0.199276
#> [279,] 15.6992 -0.009580 -0.000377 -0.014367 0.029116 0.197181
#> [280,] 8.2191 -0.081297 0.087869 -0.003100 0.024502 0.089604
#> [281,] 15.0701 -0.001933 0.015176 -0.011676 0.025438 0.192309
#> [282,] 15.9287 -0.010950 -0.000821 -0.014963 0.029845 0.191330
#> [283,] 17.9893 -0.026646 -0.041135 -0.015035 0.028410 0.232701
#> [284,] 18.8572 -0.033786 -0.055946 -0.015044 0.028694 0.238887
#> [285,] 19.5324 -0.041503 -0.064945 -0.014727 0.028271 0.235259
#> [286,] 20.7775 -0.053434 -0.085184 -0.009257 0.020046 0.236640
#> [287,] 16.9735 -0.032895 -0.065666 -0.001117 0.022132 0.238103
#> [288,] 16.5340 -0.030907 -0.061317 -0.001324 0.022882 0.237518
#> [289,] 20.8375 -0.053335 -0.079394 -0.013449 0.025312 0.229815
#> [290,] 18.1113 -0.028366 -0.046264 -0.015021 0.028745 0.235873
#> [291,] 17.8601 -0.025731 -0.039360 -0.014969 0.028332 0.232263
#> [292,] 16.5654 -0.029566 -0.061476 -0.002022 0.024028 0.236988
#> [293,] 17.0192 -0.035406 -0.059301 -0.010721 0.027714 0.212950
#> [294,] 20.7864 -0.043782 -0.112079 -0.020800 0.027329 0.195748
#> [295,] 9.0656 -0.040920 0.013650 -0.004245 -0.009114 0.262027
#> [296,] 7.9927 -0.041941 0.026504 -0.001909 -0.013255 0.267773
#> [297,] 7.4678 -0.041873 0.030278 -0.000926 -0.015414 0.276754
#> [298,] 7.0006 -0.041983 0.034746 0.000134 -0.017312 0.282317
#> [299,] 6.4706 -0.047249 0.048774 0.004787 -0.024116 0.262992
#> [300,] 8.3739 -0.045604 0.023701 -0.001110 -0.014279 0.252148
#> [301,] 9.6611 -0.045162 0.006653 -0.003565 -0.009480 0.241605
#> [302,] 13.8378 -0.042986 -0.042357 -0.011446 0.005110 0.220539
#> [303,] 16.2945 -0.044870 -0.065736 -0.015339 0.011417 0.210648
#> [304,] 19.7412 -0.049379 -0.097054 -0.020099 0.018673 0.210029
#> [305,] 21.2780 -0.046690 -0.110816 -0.020845 0.024982 0.217469
#> [306,] 11.0234 -0.039439 -0.007324 -0.008049 -0.001768 0.248556
#> [307,] 20.9300 -0.042843 -0.104830 -0.018921 0.027110 0.223513
#> [308,] 19.1415 -0.037531 -0.086631 -0.015261 0.029601 0.209585
#> [309,] 16.1930 -0.034344 -0.048465 -0.008214 0.026761 0.220708
#> [310,] 18.3754 -0.047952 -0.085714 -0.018283 0.015593 0.206673
#> [311,] 21.1639 -0.046742 -0.113407 -0.021327 0.024174 0.209315
#> [312,] 14.5980 -0.038429 -0.048798 -0.013441 0.009950 0.226188
#> [313,] 7.1511 -0.041626 0.033211 -0.000293 -0.016484 0.281206
#> [314,] 9.7372 -0.039709 0.004949 -0.005765 -0.006438 0.261241
#> [315,] 10.9207 -0.028992 -0.026441 -0.003034 -0.001744 0.189012
#> [316,] 10.9592 -0.024226 -0.021728 -0.002303 -0.001704 0.183915
#> [317,] 10.8074 -0.022811 -0.012345 -0.001735 -0.001188 0.174092
#> [318,] 10.9138 -0.009694 -0.019444 -0.001754 -0.002658 0.192005
#> [319,] 9.4150 -0.022056 0.018864 -0.000254 0.001828 0.150049
#> [320,] 10.8892 -0.006233 0.021680 -0.006208 -0.007595 0.203318
#> [321,] 8.3080 -0.028512 0.056289 0.002712 0.002673 0.116725
#> [322,] 6.7250 -0.012644 0.113167 0.013405 0.001857 0.153725
#> [323,] 8.9663 -0.024055 0.031326 0.000192 0.002138 0.141632
#> [324,] 8.5853 -0.023581 0.063649 0.005000 -0.000155 0.120152
#> [325,] 10.2770 -0.009682 0.012542 -0.001509 -0.002751 0.174750
#> [326,] 10.8844 -0.006391 -0.013337 -0.001415 -0.002730 0.188063
#> [327,] 10.9150 -0.009581 -0.019136 -0.001702 -0.002625 0.191424
#> [328,] 10.7220 -0.016722 -0.028393 -0.002630 -0.002323 0.200010
#> [329,] 7.8126 -0.108667 0.086354 0.006615 0.034922 0.080849
#> [330,] 7.8131 -0.108654 0.086391 0.006613 0.034920 0.080802
#> [331,] 10.1279 -0.018501 0.002862 -0.000864 -0.000232 0.166661
#> [332,] 7.0651 0.015433 0.106099 0.013866 0.001524 0.152508
#> [333,] 6.8872 0.017092 0.120311 0.011672 0.006462 0.135843
#> [334,] 9.6365 -0.016306 0.091556 -0.006214 -0.008631 0.182133
#> [335,] 9.8921 -0.016710 0.056162 0.000263 -0.005478 0.160419
#> [336,] 10.3408 0.030992 0.042401 0.016947 -0.007002 0.160807
#> [337,] 7.3760 -0.023195 0.095656 0.011441 -0.000809 0.125348
#> [338,] 7.4594 0.027743 0.086025 0.015848 -0.002466 0.153123
#> [339,] 8.0986 -0.115870 0.064143 0.005821 0.033817 0.103537
#> [340,] 6.6697 0.020112 0.110928 0.013265 0.001806 0.148288
#> [341,] 6.7261 -0.001532 0.122718 0.011721 0.006675 0.148421
#> [342,] 7.4331 0.000116 0.092622 0.016153 -0.002777 0.159838
#> [343,] 3.1787 0.068672 0.121942 0.014231 -0.007444 0.149711
#> [344,] 7.7268 -0.108344 0.080013 0.007023 0.034904 0.088088
#> [345,] 10.7762 0.019255 0.018712 -0.017428 -0.006545 0.200083
#> [346,] 5.8878 0.043567 0.103573 0.014007 -0.002250 0.151117
#> [347,] 5.9463 0.035765 0.107294 0.013577 -0.001396 0.151992
#> [348,] 5.9623 0.029208 0.109170 0.013841 -0.001389 0.153024
#> [349,] 6.0349 0.028364 0.110627 0.013233 -0.000652 0.156624
#> [350,] 6.6374 0.023117 0.106522 0.013603 0.000149 0.153389
#> [351,] 6.2500 0.025635 0.118656 0.012019 0.003203 0.144638
#> [352,] 3.9483 0.058739 0.117843 0.013313 -0.005292 0.169071
#> [353,] 6.2055 0.027452 0.112840 0.012573 0.000913 0.154774
#> [354,] 5.1134 0.054778 0.107394 0.014638 -0.004270 0.147467
#> [355,] 4.3792 0.059819 0.113837 0.014375 -0.004891 0.149625
#> [356,] 6.7014 0.029269 0.117681 0.010661 0.004279 0.150522
#> [357,] 7.5851 0.025746 0.122950 0.009635 0.011164 0.129623
#> [358,] 9.0121 0.039036 0.133009 0.006804 0.012300 0.107978
#> [359,] 13.3372 0.056259 0.101416 0.009351 0.002776 0.171283
#> [360,] 15.6520 0.064859 0.097955 0.006078 -0.007475 0.229606
#> [361,] 7.4158 0.030918 0.121410 0.009285 0.006022 0.149775
#> [362,] 8.3156 0.036487 0.119603 0.009011 0.012531 0.130342
#> [363,] 7.9319 0.028784 0.120035 0.009539 0.004932 0.155088
#> [364,] 9.2180 0.026747 0.138573 0.005520 0.007451 0.116667
#> [365,] 12.5384 0.033260 0.122298 0.002828 0.000741 0.151811
#> [366,] 14.1560 0.086748 0.123534 0.002077 -0.006774 0.245979
#> [367,] 14.8839 0.073253 0.108115 0.005267 -0.006500 0.232959
#> [368,] 15.3367 0.073075 0.113167 0.002935 -0.010132 0.251301
#> [369,] 16.9887 0.072482 0.114422 -0.001910 -0.018453 0.292182
#> [370,] 17.4512 0.073959 0.109480 -0.000671 -0.017760 0.285895
#> [371,] -1.8376 0.075908 0.153834 0.015022 -0.013754 0.137732
#> [372,] 1.5134 0.066325 0.109710 0.012714 0.002502 0.135079
#> [373,] 6.3330 0.069136 0.049056 0.009263 0.019645 0.153510
#> [374,] 7.3021 0.056250 0.042009 0.007773 0.024158 0.146661
#> [375,] 1.3220 0.069922 0.112236 0.012720 0.000965 0.139164
#> [376,] 4.0357 0.062739 0.080323 0.010658 0.011892 0.136167
#> [377,] -1.7899 0.079307 0.154143 0.015965 -0.018789 0.140577
#> [378,] 5.6655 0.062665 0.060687 0.009507 0.017298 0.142327
#> [379,] 2.2273 0.063018 0.101565 0.012207 0.005487 0.132057
#> [380,] -2.6126 0.075858 0.164450 0.016454 -0.016400 0.128246
#> [381,] -0.0419 0.066564 0.128866 0.013978 -0.003449 0.131525
#> [382,] 3.5544 0.065428 0.085371 0.011011 0.010087 0.138338
#> [383,] 4.4747 0.057880 0.076436 0.010436 0.013474 0.132558
#> [384,] 3.2288 0.064021 0.089295 0.011360 0.009123 0.136240
#> [385,] -0.7552 0.070845 0.138214 0.014365 -0.007279 0.135491
#> [386,] -0.2871 0.020736 0.145415 0.034220 0.002111 0.053730
#> [387,] 1.5248 0.030577 0.115199 0.034703 0.000226 0.023249
#> [388,] 13.2569 -0.013427 0.005610 0.020674 0.001907 0.085665
#> [389,] 13.6858 0.001140 -0.002352 0.012659 0.007459 0.091624
#> [390,] 13.8571 -0.005098 0.017684 0.004678 0.001575 0.184046
#> [391,] 9.8071 0.004278 0.014665 0.018379 -0.010788 0.153573
#> [392,] 7.2778 0.014492 0.036700 0.021896 -0.012253 0.071730
#> [393,] 10.6235 0.004474 0.000612 0.020376 -0.005125 0.147853
#> [394,] 11.1123 -0.009803 0.015301 0.022686 0.004169 0.112400
#> [395,] 13.2758 0.012918 0.003052 0.007007 0.010543 0.104906
#> [396,] -0.2294 0.022464 0.143875 0.034151 0.001896 0.048504
#> [397,] 12.9059 -0.005725 0.011735 0.016364 0.007726 0.073140
#> [398,] 13.1365 0.003411 0.013164 0.004756 0.008076 0.152534
#> [399,] 20.0304 -0.089998 -0.070927 0.004178 -0.033329 0.261047
#> [400,] 17.0802 -0.051227 -0.071810 0.016903 -0.014106 0.236652
#> [401,] 16.8570 -0.039862 -0.076647 0.017947 -0.012142 0.248082
#> [402,] 15.8482 -0.019765 -0.076258 0.021317 -0.008295 0.256822
#> [403,] 15.0414 -0.011477 -0.069548 0.023509 -0.006393 0.250218
#> [404,] 15.3298 -0.017266 -0.071113 0.022720 -0.007425 0.248101
#> [405,] 16.1335 -0.039812 -0.067312 0.020866 -0.011146 0.228274
#> [406,] 14.2581 -0.005762 -0.061775 0.026753 -0.005453 0.234060
#> [407,] 14.0237 -0.001757 -0.059195 0.027317 -0.004887 0.230161
#> [408,] 13.6876 0.006912 -0.053684 0.027585 -0.003687 0.217653
#> [409,] 13.6601 0.023030 -0.005657 0.008912 0.010226 0.070474
#> [410,] 13.9285 0.005166 -0.054370 0.027294 -0.004209 0.210932
#> [411,] 13.8232 0.012054 -0.047349 0.025838 -0.002753 0.190495
#> [412,] 13.6492 0.018983 -0.041685 0.024002 -0.000866 0.177784
#> [413,] 13.3146 0.019221 -0.041354 0.024336 -0.000511 0.192346
#> [414,] 13.3694 0.021791 -0.036236 0.021868 0.000813 0.177954
#> [415,] 13.8791 0.024733 -0.030966 0.018767 0.002484 0.135353
#> [416,] 13.3138 0.026954 -0.025614 0.016652 0.004114 0.152760
#> [417,] 13.4336 0.026787 -0.010111 0.010653 0.009122 0.089495
#> [418,] 14.4861 0.002511 -0.011806 0.017390 0.003188 0.062684
#> [419,] 13.3670 0.027865 -0.021189 0.014750 0.005418 0.136428
#> [420,] 14.2046 -0.003033 -0.063892 0.026348 -0.004982 0.243808
#> [421,] 13.3765 0.015638 -0.045739 0.025973 -0.001692 0.202621
#> [422,] 13.7805 0.028071 -0.024028 0.015661 0.005031 0.116092
#> [423,] 13.7239 -0.124661 0.096777 0.015685 -0.063903 0.066960
#> [424,] 14.2961 -0.130054 0.094556 0.006358 -0.056861 0.085721
#> [425,] 14.2711 -0.111981 0.096203 0.004651 -0.049136 0.071669
#> [426,] 15.2781 -0.092646 0.028500 0.010799 -0.029004 0.095606
#> [427,] 13.8592 -0.120749 0.109839 0.005448 -0.055957 0.069356
#> [428,] 15.5069 -0.115815 0.038578 0.006441 -0.036332 0.121023
#> [429,] 10.9412 -0.101439 0.116870 0.026383 -0.059399 0.021114
#> [430,] 16.6315 -0.000985 -0.035087 0.016765 -0.001127 0.048045
#> [431,] 10.6486 -0.114336 0.147514 0.013604 -0.054830 0.027807
#> [432,] 15.3006 -0.089885 0.043310 0.010421 -0.030070 0.087360
#> [433,] 14.8000 -0.106093 0.063410 0.007019 -0.038450 0.080373
#> [434,] 7.7351 -0.093335 0.186010 0.014333 -0.046894 0.014725
#> [435,] 14.7150 -0.127755 0.079788 0.004974 -0.050697 0.098396
#> [436,] 14.9611 -0.129611 0.070515 0.007581 -0.055812 0.115611
#> [437,] 13.9181 -0.117526 0.076466 0.016070 -0.061478 0.103291
#> [438,] 13.3962 -0.114586 0.115311 0.006236 -0.053293 0.064914
#> [439,] 14.1916 -0.121299 0.077067 0.013643 -0.060105 0.104283
#> [440,] 2.7745 0.047423 0.089981 0.022713 -0.012944 0.092932
#> [441,] 0.0978 0.061728 0.126917 0.020164 -0.015490 0.106570
#> [442,] 5.9560 0.028733 0.046428 0.025382 -0.007242 0.074488
#> [443,] -1.6690 0.074022 0.151426 0.017515 -0.017352 0.126116
#> [444,] 1.5024 0.057727 0.106375 0.020177 -0.016204 0.114398
#> [445,] -0.8328 0.067717 0.139750 0.018903 -0.016298 0.114979
#> [446,] 12.1677 0.020134 -0.023452 0.019427 0.004611 0.172303
#> [447,] 12.7479 0.023041 -0.031646 0.020695 0.002646 0.184912
#> [448,] 12.7001 0.025000 -0.020133 0.015142 0.005618 0.163755
#> [449,] 11.5395 0.020109 -0.008779 0.013808 0.008309 0.164426
#> [450,] 11.8335 0.013183 0.020993 0.002487 0.011497 0.146680
#> [451,] 11.9663 0.022479 0.002325 0.008565 0.011670 0.121265
#> [452,] 9.3917 -0.019843 0.094326 -0.001788 0.004661 0.007692
#> [453,] 10.6076 -0.030959 0.081448 -0.003915 0.008293 0.015423
#> [454,] 14.1877 0.014583 0.054355 -0.006773 0.015428 0.049119
#> [455,] 10.7405 -0.018463 0.075896 -0.005840 0.012551 0.091447
#> [456,] 10.4255 -0.053902 0.081762 -0.003098 0.004247 0.010592
#> [457,] 14.2715 0.012849 0.067653 -0.012322 0.014671 0.058237
#> [458,] 14.0262 0.004979 0.041413 -0.003508 0.015297 0.046279
#> [459,] 5.4373 -0.038845 0.068460 0.012214 -0.033182 0.246053
#> [460,] 11.3987 -0.016932 0.072467 -0.007240 0.013245 0.090600
#> [461,] 14.2101 0.005664 0.070650 -0.011216 0.009978 0.062953
#> [462,] 10.7020 -0.030988 0.080539 -0.004009 0.008466 0.015597
#> [463,] 8.6639 0.030405 0.054254 0.002944 0.021653 0.110141
#> [464,] 11.1036 0.029610 0.044636 -0.002091 0.024160 0.099192
#> [465,] 2.8712 -0.033651 0.133868 0.012391 0.002837 0.013070
#> [466,] 11.8407 0.014200 0.052115 -0.002673 0.016078 0.081768
#> [467,] 9.1827 0.031810 0.049739 0.001577 0.023067 0.113364
#> [468,] 10.7043 0.032302 0.041733 -0.001348 0.025688 0.108726
#> [469,] 9.3357 0.037496 0.041162 0.000782 0.027318 0.127084
#> [470,] 1.0508 -0.027770 0.167157 0.018304 -0.005230 -0.029208
#> [471,] 8.5807 0.026503 0.057129 0.003801 0.019540 0.103041
#> [472,] 8.7586 0.025748 0.056497 0.003561 0.019317 0.102070
#> [473,] 14.4803 -0.016352 0.033061 -0.004296 0.019048 0.020020
#> [474,] 12.5163 -0.040538 0.065474 -0.006485 0.009525 0.016951
#> [475,] 13.3015 -0.045452 0.059634 -0.007657 0.009383 0.017059
#> [476,] 14.9721 -0.029274 0.032729 -0.007177 0.016973 0.024264
#> [477,] 15.8214 -0.036688 0.027435 -0.009178 0.016024 0.024207
#> [478,] 15.1726 -0.002700 0.019059 -0.004509 0.022126 0.032702
#> [479,] 16.0964 0.012296 0.001071 -0.003648 0.024643 0.062999
#> [480,] 16.0633 0.021308 0.009463 -0.003841 0.024179 0.070190
#> [481,] 15.9940 0.003085 0.004552 -0.005167 0.024589 0.044866
#> [482,] 13.0105 -0.060023 0.077743 -0.010494 0.030072 -0.066970
#> [483,] 15.5330 -0.036042 0.030568 -0.008862 0.015717 0.023762
#> [484,] 16.0698 -0.034399 0.022583 -0.008848 0.017325 0.025479
#> [485,] 16.4516 0.017720 -0.002007 -0.003487 0.025309 0.072957
#> [486,] 10.3406 -0.059186 0.077729 -0.000209 0.002034 0.008855
#> [487,] 11.5789 -0.058968 0.072016 -0.003778 0.003367 0.010912
#> [488,] 11.7675 -0.059182 0.070770 -0.004029 0.003284 0.011048
#> [489,] 12.1489 -0.055128 0.070442 -0.006024 0.005191 0.012809
#> [490,] 13.1472 -0.055251 0.064697 -0.008118 0.005926 0.014249
#> [491,] 13.6280 -0.053896 0.061523 -0.009147 0.006799 0.015429
#> [492,] 11.8927 -0.056904 0.071526 -0.005223 0.004462 0.011986
#> [493,] 11.6693 -0.058160 0.072229 -0.004408 0.003859 0.011320
#> [494,] 11.5218 -0.058286 0.072997 -0.004057 0.003725 0.011078
#> [495,] 11.0790 -0.059288 0.074003 -0.002206 0.002727 0.010018
#> [496,] 10.7330 -0.059349 0.076042 -0.001500 0.002525 0.009526
#> [497,] 11.2673 -0.059054 0.073683 -0.003066 0.003158 0.010449
#> [498,] 11.4014 -0.059170 0.072647 -0.003180 0.003098 0.010567
#> [499,] 11.3537 -0.059255 0.072627 -0.002899 0.002941 0.010435
#> [500,] 11.7660 -0.058802 0.071223 -0.004316 0.003579 0.011231
#> [501,] 12.2935 -0.058683 0.068697 -0.005633 0.003915 0.011982
#> [502,] 11.6144 -0.059269 0.071307 -0.003515 0.003078 0.010777
#> [503,] 12.3144 -0.056905 0.069215 -0.006131 0.004723 0.012575
#> [504,] 14.0737 -0.052209 0.057481 -0.009841 0.007895 0.016636
#> [505,] 12.2239 -0.058204 0.069285 -0.005633 0.004140 0.012052
#> [506,] 11.4811 -0.058875 0.072690 -0.003648 0.003389 0.010817
#> [507,] 11.6215 -0.058802 0.071978 -0.003990 0.003502 0.011038
#> [508,] 12.1830 -0.058579 0.069303 -0.005409 0.003927 0.011864
#> [509,] 11.5938 -0.059113 0.071741 -0.003683 0.003257 0.010858
#> [510,] 11.7179 -0.058397 0.071792 -0.004410 0.003773 0.011313
#> [511,] 11.8235 -0.056236 0.072129 -0.005211 0.004637 0.012062
#> [512,] 11.9719 -0.057344 0.070922 -0.005288 0.004351 0.011979
#> [513,] 11.7319 -0.057267 0.072321 -0.004811 0.004242 0.011663
#> [514,] 11.8176 -0.059270 0.070368 -0.004047 0.003206 0.011054
#> X9
#> [1,] -2.33e-02
#> [2,] -3.66e-02
#> [3,] -3.31e-02
#> [4,] -3.74e-02
#> [5,] 1.94e-03
#> [6,] -1.11e-02
#> [7,] -3.00e-03
#> [8,] 4.69e-03
#> [9,] 1.52e-03
#> [10,] -4.91e-03
#> [11,] -1.01e-02
#> [12,] -3.41e-02
#> [13,] -2.66e-02
#> [14,] -3.35e-02
#> [15,] -9.20e-03
#> [16,] 1.89e-03
#> [17,] -9.90e-03
#> [18,] 2.09e-04
#> [19,] 5.45e-03
#> [20,] 6.16e-03
#> [21,] -3.00e-02
#> [22,] -7.27e-03
#> [23,] -3.91e-02
#> [24,] -7.36e-02
#> [25,] -7.53e-02
#> [26,] -7.45e-02
#> [27,] -5.31e-02
#> [28,] -6.08e-02
#> [29,] -6.06e-02
#> [30,] -7.73e-02
#> [31,] -6.05e-02
#> [32,] -5.46e-02
#> [33,] -4.15e-02
#> [34,] -4.21e-02
#> [35,] -4.47e-02
#> [36,] -4.04e-02
#> [37,] -2.41e-02
#> [38,] -4.84e-02
#> [39,] -4.21e-02
#> [40,] -5.09e-02
#> [41,] -4.84e-02
#> [42,] -5.54e-02
#> [43,] -7.92e-02
#> [44,] -1.01e-01
#> [45,] -8.46e-02
#> [46,] -7.28e-02
#> [47,] -2.58e-02
#> [48,] -2.56e-02
#> [49,] -5.67e-02
#> [50,] -6.18e-02
#> [51,] -5.18e-02
#> [52,] -4.96e-02
#> [53,] -4.46e-02
#> [54,] -4.34e-02
#> [55,] -7.24e-02
#> [56,] -2.88e-02
#> [57,] -6.59e-02
#> [58,] -8.43e-02
#> [59,] -1.24e-01
#> [60,] -1.31e-01
#> [61,] -1.30e-01
#> [62,] -1.35e-01
#> [63,] -1.42e-01
#> [64,] -5.43e-02
#> [65,] -1.10e-01
#> [66,] -9.19e-02
#> [67,] -9.37e-02
#> [68,] -9.76e-02
#> [69,] -1.23e-01
#> [70,] -1.25e-01
#> [71,] -1.27e-01
#> [72,] -1.37e-01
#> [73,] -1.32e-01
#> [74,] -1.36e-01
#> [75,] -1.35e-01
#> [76,] -1.21e-01
#> [77,] -7.28e-02
#> [78,] 3.31e-02
#> [79,] -7.10e-02
#> [80,] -3.58e-02
#> [81,] -1.28e-01
#> [82,] -1.03e-01
#> [83,] -3.74e-02
#> [84,] -9.91e-02
#> [85,] 1.83e-02
#> [86,] -1.26e-01
#> [87,] -9.59e-02
#> [88,] -7.27e-02
#> [89,] -3.81e-02
#> [90,] -7.95e-02
#> [91,] -8.72e-02
#> [92,] -7.20e-02
#> [93,] -7.24e-03
#> [94,] 7.19e-03
#> [95,] -6.53e-02
#> [96,] -5.62e-02
#> [97,] -9.35e-02
#> [98,] -4.90e-02
#> [99,] -4.24e-02
#> [100,] -3.48e-02
#> [101,] -6.61e-02
#> [102,] -6.21e-02
#> [103,] -1.07e-01
#> [104,] -1.57e-01
#> [105,] -1.11e-01
#> [106,] -2.41e-02
#> [107,] -3.39e-02
#> [108,] -7.49e-02
#> [109,] -6.39e-02
#> [110,] -1.08e-01
#> [111,] -1.01e-01
#> [112,] -9.17e-02
#> [113,] -9.90e-02
#> [114,] -4.91e-02
#> [115,] -8.15e-02
#> [116,] -3.45e-02
#> [117,] -6.65e-02
#> [118,] -5.92e-02
#> [119,] -2.50e-02
#> [120,] -4.35e-02
#> [121,] -4.11e-02
#> [122,] -1.13e-01
#> [123,] -5.98e-02
#> [124,] -5.32e-02
#> [125,] -4.82e-02
#> [126,] -1.15e-02
#> [127,] -1.41e-02
#> [128,] -2.92e-02
#> [129,] -2.93e-02
#> [130,] -1.48e-02
#> [131,] -1.49e-02
#> [132,] -2.21e-02
#> [133,] -1.06e-02
#> [134,] -1.97e-02
#> [135,] -1.62e-02
#> [136,] -1.59e-02
#> [137,] -1.69e-02
#> [138,] -1.06e-02
#> [139,] -2.37e-02
#> [140,] -1.85e-02
#> [141,] -2.59e-02
#> [142,] 7.07e-03
#> [143,] -3.36e-02
#> [144,] -3.96e-02
#> [145,] -4.40e-02
#> [146,] 2.16e-02
#> [147,] -3.56e-02
#> [148,] 6.21e-02
#> [149,] 1.03e-01
#> [150,] 6.72e-02
#> [151,] 1.08e-01
#> [152,] 7.39e-02
#> [153,] 9.14e-02
#> [154,] 1.05e-01
#> [155,] -4.28e-02
#> [156,] -2.87e-02
#> [157,] -3.08e-02
#> [158,] -3.19e-02
#> [159,] -2.66e-02
#> [160,] -3.51e-02
#> [161,] -1.19e-02
#> [162,] 1.29e-02
#> [163,] -9.43e-03
#> [164,] 5.99e-03
#> [165,] 5.04e-02
#> [166,] 8.95e-02
#> [167,] 6.26e-02
#> [168,] 6.38e-02
#> [169,] 6.76e-02
#> [170,] 6.08e-02
#> [171,] 1.27e-02
#> [172,] 1.49e-03
#> [173,] -2.30e-02
#> [174,] -2.90e-02
#> [175,] -3.15e-02
#> [176,] -2.67e-02
#> [177,] -9.38e-03
#> [178,] 3.43e-02
#> [179,] -1.34e-02
#> [180,] -4.00e-03
#> [181,] 3.63e-04
#> [182,] 6.27e-02
#> [183,] -3.09e-02
#> [184,] -2.07e-02
#> [185,] -5.26e-03
#> [186,] 8.41e-02
#> [187,] 4.26e-02
#> [188,] -1.60e-02
#> [189,] -3.15e-02
#> [190,] -3.07e-02
#> [191,] -3.86e-02
#> [192,] -3.54e-02
#> [193,] -4.41e-02
#> [194,] -4.29e-02
#> [195,] -2.76e-02
#> [196,] 2.80e-03
#> [197,] 4.32e-03
#> [198,] 1.36e-02
#> [199,] -5.86e-05
#> [200,] 1.40e-02
#> [201,] 1.91e-02
#> [202,] 2.80e-02
#> [203,] 6.50e-02
#> [204,] 7.50e-02
#> [205,] 4.32e-02
#> [206,] 2.49e-02
#> [207,] 1.20e-02
#> [208,] 1.28e-02
#> [209,] -2.00e-02
#> [210,] -3.31e-02
#> [211,] -3.13e-02
#> [212,] -3.57e-02
#> [213,] -3.55e-02
#> [214,] -2.41e-02
#> [215,] -9.01e-04
#> [216,] 1.79e-02
#> [217,] -3.14e-02
#> [218,] 1.54e-02
#> [219,] -1.24e-03
#> [220,] -8.56e-03
#> [221,] -3.53e-02
#> [222,] 3.18e-04
#> [223,] -4.41e-02
#> [224,] -2.29e-02
#> [225,] 1.63e-03
#> [226,] -2.23e-02
#> [227,] -1.87e-02
#> [228,] -1.14e-02
#> [229,] -2.19e-02
#> [230,] -2.65e-02
#> [231,] -3.23e-02
#> [232,] -6.82e-02
#> [233,] -4.62e-02
#> [234,] -8.08e-02
#> [235,] -7.65e-02
#> [236,] -7.37e-02
#> [237,] -6.72e-02
#> [238,] -7.05e-02
#> [239,] -6.76e-02
#> [240,] -6.51e-02
#> [241,] -6.60e-02
#> [242,] -5.49e-02
#> [243,] -5.57e-02
#> [244,] -2.71e-02
#> [245,] -1.33e-02
#> [246,] -4.48e-03
#> [247,] -7.09e-03
#> [248,] 1.31e-02
#> [249,] 2.78e-02
#> [250,] 3.12e-02
#> [251,] -1.84e-02
#> [252,] -3.70e-02
#> [253,] -3.62e-02
#> [254,] -4.14e-02
#> [255,] -4.85e-02
#> [256,] -5.32e-02
#> [257,] -3.17e-02
#> [258,] -5.92e-02
#> [259,] -8.10e-02
#> [260,] -6.71e-02
#> [261,] -6.19e-02
#> [262,] -4.73e-02
#> [263,] -8.42e-03
#> [264,] -4.47e-02
#> [265,] -7.45e-02
#> [266,] -1.27e-02
#> [267,] -2.01e-03
#> [268,] -2.03e-02
#> [269,] -2.07e-02
#> [270,] -2.75e-02
#> [271,] -2.75e-02
#> [272,] -2.10e-02
#> [273,] -2.28e-02
#> [274,] -6.30e-02
#> [275,] -5.43e-02
#> [276,] -5.30e-02
#> [277,] -4.96e-02
#> [278,] -4.83e-02
#> [279,] -4.93e-02
#> [280,] -2.12e-02
#> [281,] -5.45e-02
#> [282,] -5.16e-02
#> [283,] -4.03e-02
#> [284,] -3.80e-02
#> [285,] -3.75e-02
#> [286,] -3.09e-02
#> [287,] -1.57e-02
#> [288,] -1.52e-02
#> [289,] -3.78e-02
#> [290,] -3.76e-02
#> [291,] -4.03e-02
#> [292,] -1.67e-02
#> [293,] -1.95e-02
#> [294,] -1.06e-02
#> [295,] 5.48e-02
#> [296,] 6.01e-02
#> [297,] 6.48e-02
#> [298,] 6.80e-02
#> [299,] 6.79e-02
#> [300,] 5.95e-02
#> [301,] 5.46e-02
#> [302,] 3.39e-02
#> [303,] 1.96e-02
#> [304,] -5.98e-04
#> [305,] -1.59e-02
#> [306,] 4.28e-02
#> [307,] -2.18e-02
#> [308,] -2.09e-02
#> [309,] -1.96e-02
#> [310,] 8.83e-03
#> [311,] -1.07e-02
#> [312,] 2.47e-02
#> [313,] 6.68e-02
#> [314,] 5.17e-02
#> [315,] 6.51e-02
#> [316,] 5.84e-02
#> [317,] 5.00e-02
#> [318,] 5.48e-02
#> [319,] 3.44e-02
#> [320,] 2.09e-02
#> [321,] 1.12e-02
#> [322,] -3.55e-02
#> [323,] 2.81e-02
#> [324,] -3.93e-04
#> [325,] 3.15e-02
#> [326,] 4.84e-02
#> [327,] 5.44e-02
#> [328,] 6.83e-02
#> [329,] -3.15e-02
#> [330,] -3.16e-02
#> [331,] 4.19e-02
#> [332,] -3.53e-02
#> [333,] -4.85e-02
#> [334,] -2.87e-02
#> [335,] -4.96e-03
#> [336,] -1.76e-02
#> [337,] -2.08e-02
#> [338,] -2.08e-02
#> [339,] -1.19e-02
#> [340,] -3.39e-02
#> [341,] -4.85e-02
#> [342,] -2.50e-02
#> [343,] 9.97e-03
#> [344,] -2.47e-02
#> [345,] 4.23e-02
#> [346,] -1.44e-02
#> [347,] -1.85e-02
#> [348,] -2.03e-02
#> [349,] -2.28e-02
#> [350,] -2.83e-02
#> [351,] -3.56e-02
#> [352,] 1.58e-03
#> [353,] -2.82e-02
#> [354,] -6.24e-03
#> [355,] -1.52e-03
#> [356,] -4.12e-02
#> [357,] -6.44e-02
#> [358,] -9.21e-02
#> [359,] -1.23e-01
#> [360,] -1.44e-01
#> [361,] -5.53e-02
#> [362,] -7.34e-02
#> [363,] -6.05e-02
#> [364,] -9.29e-02
#> [365,] -1.17e-01
#> [366,] -1.49e-01
#> [367,] -1.45e-01
#> [368,] -1.51e-01
#> [369,] -1.65e-01
#> [370,] -1.68e-01
#> [371,] 4.84e-02
#> [372,] 2.44e-02
#> [373,] -8.66e-03
#> [374,] -1.88e-02
#> [375,] 2.60e-02
#> [376,] 5.82e-03
#> [377,] 5.31e-02
#> [378,] -5.67e-03
#> [379,] 1.90e-02
#> [380,] 5.22e-02
#> [381,] 3.54e-02
#> [382,] 9.55e-03
#> [383,] 2.03e-03
#> [384,] 1.18e-02
#> [385,] 4.06e-02
#> [386,] 6.08e-03
#> [387,] 1.43e-02
#> [388,] -2.62e-02
#> [389,] -2.39e-02
#> [390,] -3.29e-02
#> [391,] 2.75e-02
#> [392,] 4.16e-02
#> [393,] 2.01e-02
#> [394,] -1.12e-02
#> [395,] -2.32e-02
#> [396,] 7.23e-03
#> [397,] -2.92e-02
#> [398,] -2.61e-02
#> [399,] 1.82e-02
#> [400,] 1.57e-02
#> [401,] 1.78e-02
#> [402,] 1.94e-02
#> [403,] 1.82e-02
#> [404,] 1.89e-02
#> [405,] 1.48e-02
#> [406,] 1.63e-02
#> [407,] 1.53e-02
#> [408,] 1.22e-02
#> [409,] -2.22e-02
#> [410,] 1.11e-02
#> [411,] 5.80e-03
#> [412,] 2.34e-03
#> [413,] 5.12e-03
#> [414,] 1.18e-03
#> [415,] -7.49e-03
#> [416,] -5.36e-03
#> [417,] -1.72e-02
#> [418,] -2.59e-02
#> [419,] -8.90e-03
#> [420,] 1.83e-02
#> [421,] 7.92e-03
#> [422,] -1.15e-02
#> [423,] -1.78e-02
#> [424,] -2.20e-02
#> [425,] -3.34e-02
#> [426,] -1.79e-02
#> [427,] -3.18e-02
#> [428,] -1.45e-02
#> [429,] -1.88e-02
#> [430,] -2.63e-02
#> [431,] -3.40e-02
#> [432,] -3.02e-02
#> [433,] -2.63e-02
#> [434,] -4.50e-02
#> [435,] -2.07e-02
#> [436,] -1.19e-02
#> [437,] -7.00e-03
#> [438,] -3.57e-02
#> [439,] -9.82e-03
#> [440,] 4.53e-02
#> [441,] 4.98e-02
#> [442,] 3.70e-02
#> [443,] 5.25e-02
#> [444,] 5.13e-02
#> [445,] 5.11e-02
#> [446,] 4.59e-03
#> [447,] 4.12e-03
#> [448,] -2.23e-03
#> [449,] 2.32e-03
#> [450,] -1.83e-02
#> [451,] -9.98e-03
#> [452,] -3.60e-02
#> [453,] -3.88e-02
#> [454,] -8.93e-02
#> [455,] -5.57e-02
#> [456,] -2.28e-02
#> [457,] -9.79e-02
#> [458,] -7.46e-02
#> [459,] 6.53e-02
#> [460,] -6.15e-02
#> [461,] -9.37e-02
#> [462,] -3.93e-02
#> [463,] -3.65e-02
#> [464,] -5.98e-02
#> [465,] 2.61e-03
#> [466,] -6.25e-02
#> [467,] -4.06e-02
#> [468,] -5.51e-02
#> [469,] -4.10e-02
#> [470,] 2.75e-03
#> [471,] -3.50e-02
#> [472,] -3.63e-02
#> [473,] -6.58e-02
#> [474,] -4.56e-02
#> [475,] -4.78e-02
#> [476,] -6.19e-02
#> [477,] -6.29e-02
#> [478,] -7.16e-02
#> [479,] -7.92e-02
#> [480,] -8.93e-02
#> [481,] -7.45e-02
#> [482,] -6.81e-02
#> [483,] -6.20e-02
#> [484,] -6.45e-02
#> [485,] -8.47e-02
#> [486,] -1.51e-02
#> [487,] -2.60e-02
#> [488,] -2.71e-02
#> [489,] -3.44e-02
#> [490,] -4.20e-02
#> [491,] -4.64e-02
#> [492,] -3.09e-02
#> [493,] -2.79e-02
#> [494,] -2.66e-02
#> [495,] -2.12e-02
#> [496,] -1.86e-02
#> [497,] -2.35e-02
#> [498,] -2.42e-02
#> [499,] -2.35e-02
#> [500,] -2.78e-02
#> [501,] -3.21e-02
#> [502,] -2.56e-02
#> [503,] -3.42e-02
#> [504,] -5.02e-02
#> [505,] -3.22e-02
#> [506,] -2.54e-02
#> [507,] -2.66e-02
#> [508,] -3.14e-02
#> [509,] -2.58e-02
#> [510,] -2.79e-02
#> [511,] -3.10e-02
#> [512,] -3.11e-02
#> [513,] -2.93e-02
#> [514,] -2.72e-02datgwr2 %>% mutate(beta) %>% head(5)
#> Y X1 X5 X6 X7 X8 X9 beta.Intercept beta.X1
#> 1 1648096 19.0 65.3 71.6 87.5 5.71 71.2 2.19588 -0.05938
#> 2 1780419 20.4 67.4 69.6 78.6 8.36 62.9 7.83196 -0.10411
#> 3 4345784 13.2 64.4 62.5 79.7 6.46 60.9 3.71974 -0.07743
#> 4 3487157 13.4 68.2 62.7 86.7 6.43 69.6 4.67283 -0.08745
#> 5 8433526 14.4 68.7 66.8 83.2 7.13 59.5 4.52376 -0.06140
#> beta.X5 beta.X6 beta.X7 beta.X8 beta.X9
#> 1 0.18258 -0.00309 0.01576 0.02211 -0.02334
#> 2 0.13055 -0.00591 0.01553 -0.02178 -0.03663
#> 3 0.17329 -0.00431 0.01995 -0.01210 -0.03307
#> 4 0.16616 -0.00458 0.02060 -0.02297 -0.03736
#> 5 0.12841 -0.01350 0.01555 0.12212 0.00194b <- data.frame(beta)
head(b)
#> Intercept X1 X5 X6 X7 X8 X9
#> 1 2.20 -0.0594 0.183 -0.00309 0.0158 0.0221 -0.02334
#> 2 7.83 -0.1041 0.131 -0.00591 0.0155 -0.0218 -0.03663
#> 3 3.72 -0.0774 0.173 -0.00431 0.0200 -0.0121 -0.03307
#> 4 4.67 -0.0874 0.166 -0.00458 0.0206 -0.0230 -0.03736
#> 5 4.52 -0.0614 0.128 -0.01350 0.0155 0.1221 0.00194
#> 6 4.78 -0.0706 0.134 -0.00993 0.0200 0.0683 -0.01114Ymodel <- mgwrsar_0_0_kv_gauss$Y
head(Ymodel)
#> [,1]
#> 1 14.3
#> 2 14.4
#> 3 15.3
#> 4 15.1
#> 5 15.9
#> 6 15.6# parameter
koefisien <- b %>% mutate(Y = Ymodel)
head(koefisien)
#> Intercept X1 X5 X6 X7 X8 X9 Y
#> 1 2.20 -0.0594 0.183 -0.00309 0.0158 0.0221 -0.02334 14.3
#> 2 7.83 -0.1041 0.131 -0.00591 0.0155 -0.0218 -0.03663 14.4
#> 3 3.72 -0.0774 0.173 -0.00431 0.0200 -0.0121 -0.03307 15.3
#> 4 4.67 -0.0874 0.166 -0.00458 0.0206 -0.0230 -0.03736 15.1
#> 5 4.52 -0.0614 0.128 -0.01350 0.0155 0.1221 0.00194 15.9
#> 6 4.78 -0.0706 0.134 -0.00993 0.0200 0.0683 -0.01114 15.6std_error <- mgwrsar_0_0_kv_gauss$sev
std_error2 <- data.frame(std_error)
head(std_error2)
#> Intercept X1 X5 X6 X7 X8 X9
#> 1 3.01 0.0236 0.0396 0.00638 0.0102 0.0528 0.0164
#> 2 3.25 0.0269 0.0469 0.00695 0.0107 0.0584 0.0200
#> 3 3.62 0.0307 0.0547 0.00915 0.0125 0.0612 0.0207
#> 4 3.84 0.0337 0.0596 0.00974 0.0128 0.0645 0.0221
#> 5 5.00 0.0361 0.0701 0.01131 0.0176 0.0616 0.0222
#> 6 4.57 0.0341 0.0658 0.01089 0.0162 0.0617 0.0218